 |
|
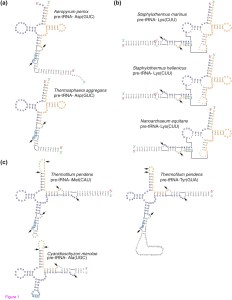
Figure 1 – Predicted secondary structures of trans-spliced and permuted precursor tRNAs
(a) Mature tRNAAsp(GUC) in A. pernix and T. aggregans are formed by joining the 5′ half and the 3′ half at position 37/38 after splicing at the bulge-helix-bulge (BHB) motif. (b) The 5′ half and the 3′ half of trans-spliced tRNALys(CUU) in S. hellenicus and S. marinus join at position 30/31, same as the previously identified split tRNALys(CUU) in N. equitans [5]. (c) Circularized permuted tRNAiMet(CAU) and tRNATyr(GUA) in T. pendens have the 3′ half located upstream of the 5′ half separated by intervening sequences represented in green. The two fragments join at position 59/60, same as the T-Ψ-C loop permuted tRNAs in the red alga C. merolae [9]. Pre-tRNAAla(UGC) in C. merolae is shown for comparison. 5′ half of tRNA transcripts are represented in blue, the 3′ halves in orange. Black arrows indicate positions of splicing. Anticodons are boxed in light blue.
|
I was woefully unaware of some of the tRNA shenanigans going on in microbes until reading this paper: Genome Biology | Abstract | Discovery of permuted and recently split transfer RNAs in Archaea from Patricia Chan, Aaron Cozen and Todd Lowe. Life is pretty weird and wacky sometimes, even in components of cells that are considered “core” parts of the machinery of life. Go figure. It is worth a read …
Author: Jonathan Eisen
I am an evolutionary biologist and a Professor at U. C. Davis. (see my lab site here). My research focuses on the origin of novelty (how new processes and functions originate). To study this I focus on sequencing and analyzing genomes of organisms, especially microbes and using phylogenomic analysis
View all posts by Jonathan Eisen


