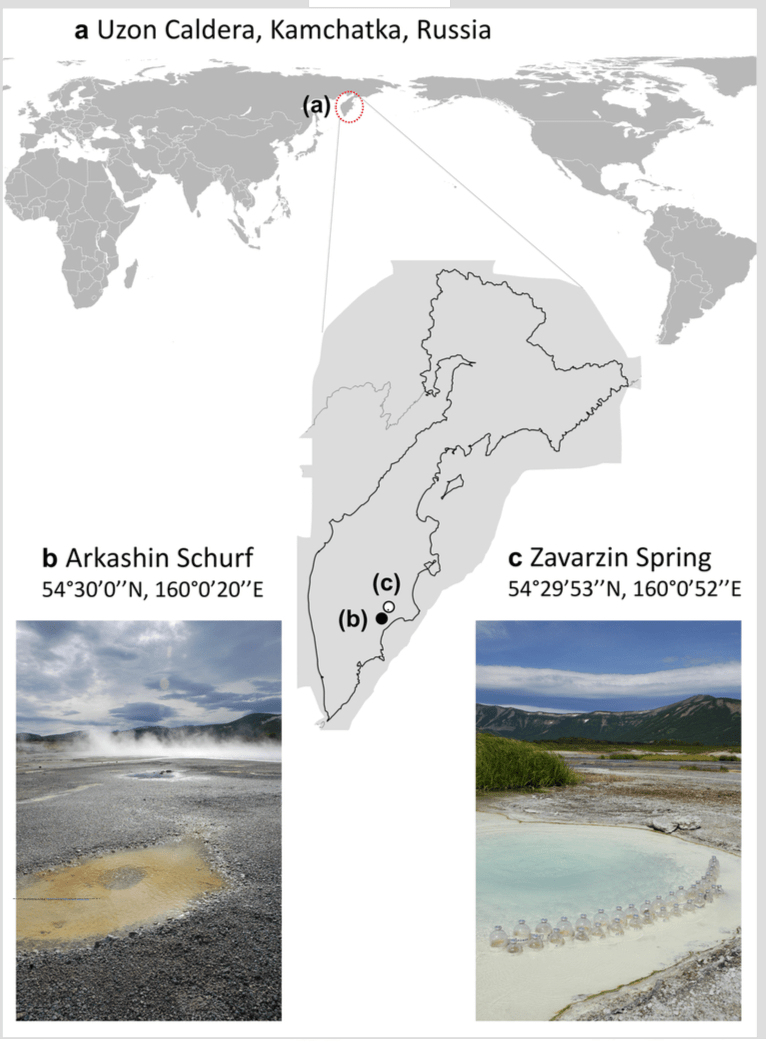
A new preprint has been posted from the lab: There and back again: metagenome-assembled genomes provide new insights into two thermal pools in Kamchatka, Russia | bioRxiv
Paper was led by
Abstract:
Culture-independent methods have contributed substantially to our understanding of global microbial diversity. Recently developed algorithms to construct whole genomes from environmental samples have further refined, corrected and revolutionized the tree of life. Here, we assembled draft metagenome-assembled genomes (MAGs) from environmental DNA extracted from two hot springs within an active volcanic ecosystem on the Kamchatka peninsula, Russia. This hydrothermal system has been intensively studied previously with regard to geochemistry, chemoautotrophy, microbial isolation, and microbial diversity. Using a shotgun metagenomics approach, we assembled population-level genomes of bacteria and archaea from two pools using DNA that had previously been characterized via 16S rRNA gene clone libraries. We recovered 36 MAGs, 29 of medium to high quality, and placed them in the context of the current microbial tree of life. We highlight MAGs representing previously underrepresented archaeal phyla (Korarchaeota, Bathyarchaeota and Aciduliprofundum) and one potentially new species within the bacterial genus Sulfurihydrogenibium. Putative functions in both pools were compared and are discussed in the context of their diverging geochemistry. This study can be considered complementary to foregoing studies in the same ecosystem as it adds more comprehensive information about phylogenetic diversity and functional potential within this highly selective habitat.
I will try to write more about it later – I am so impressed by what Laetitia and Cassie did here. This is what we call a “reboot” project in the lab – this was data that was generated and then not turned into a paper and was just sitting there. A while ago we started a “reboot” program to start to try to turn such data sets into papers and they adopted this data set. The data is Solexa shotgun metagenomic data from samples from hot springs in Kamchatka (yes, Solexa, read the paper for details). The DNA used was leftover from an old project which included a collaboration with Juergen Wiegel’s lab (and some others) that had started, at least in terms of discussions, in 2001. Anyway – I will try to write out more on the story behind this work soon. But please, check out the paper, and feel free to provide feedback.