PR from MPG here: Symbionts without borders – Bacterial partners travel the world
Authors:
- Co-First Authors: Jay T. Osvatic and Laetitia G. E. Wilkins
- Co-Last Authors: Jillian M. Petersen and Benedict Yuen
- Other authors: Lukas Leibrecht, Matthieu Leray, Sarah Zauner, Julia Polzin, Yolanda Camacho, Olivier Gros, Jan A. van Gils, Jonathan A. Eisen
Significance (from the paper)
Knowledge of host–symbiont biogeography is critical to understanding fundamental aspects of symbiosis such as host–symbiont specificity. Marine animals typically acquire their symbionts from the environment, a strategy that enables the host to associate with symbionts that are well-suited to local conditions. In contrast, we discovered that in the chemosymbiotic bivalve family Lucinidae several host species distributed across the globe are all associated with a single cosmopolitan bacterial symbiont. The genetic cohesiveness of this global symbiont species is maintained through homologous recombination across its extensive geographical range. The remarkable flexibility in the lucinid association is advantageous to both host and symbiont as it increases the likelihood of locating a compatible partner across diverse habitats spanning the globe.
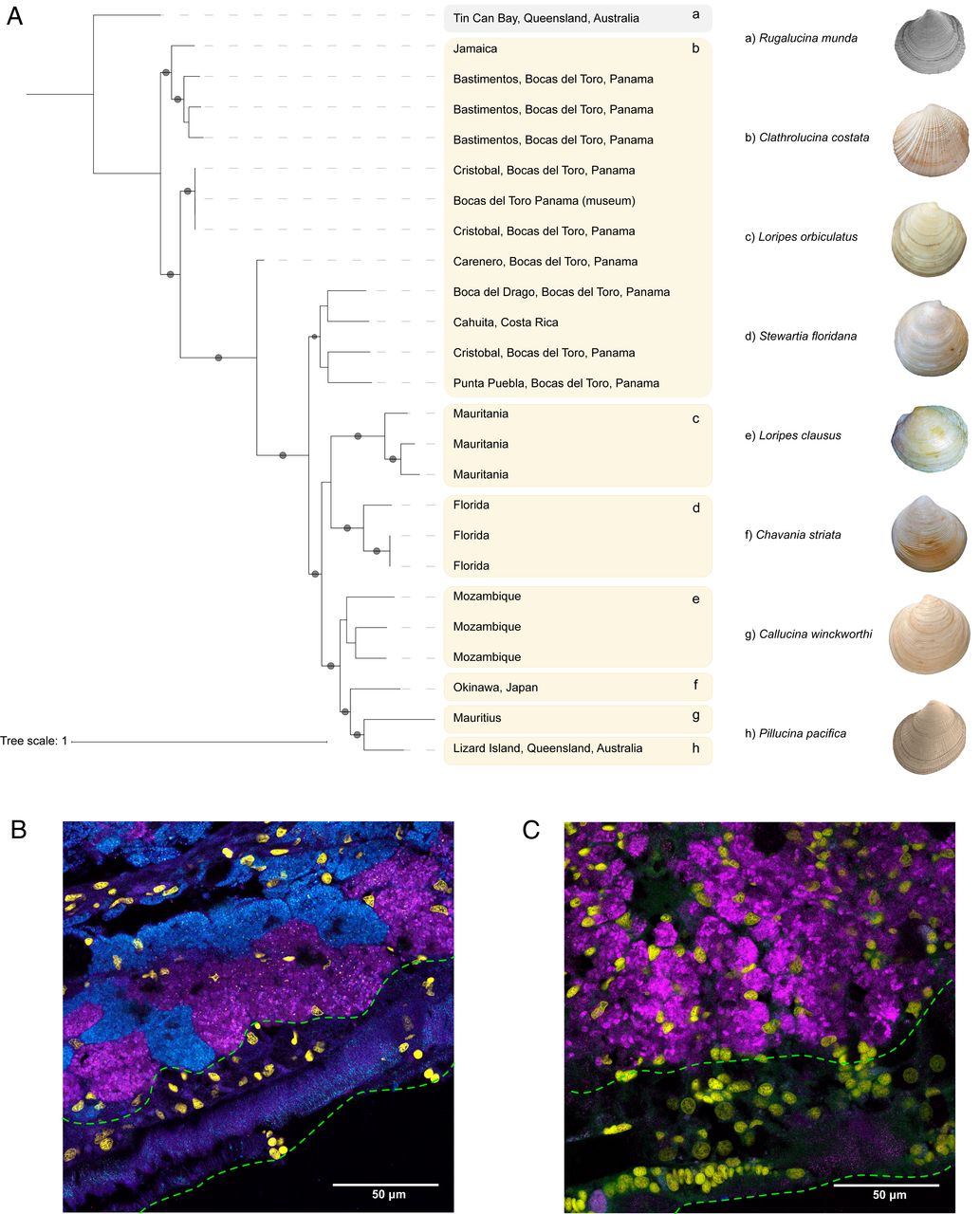
In the ocean, most hosts acquire their symbionts from the environment. Due to the immense spatial scales involved, our understanding of the biogeography of hosts and symbionts in marine systems is patchy, although this knowledge is essential for understanding fundamental aspects of symbiosis such as host–symbiont specificity and evolution. Lucinidae is the most species-rich and widely distributed family of marine bivalves hosting autotrophic bacterial endosymbionts. Previous molecular surveys identified location-specific symbiont types that “promiscuously” form associations with multiple divergent cooccurring host species. This flexibility of host–microbe pairings is thought to underpin their global success, as it allows hosts to form associations with locally adapted symbionts. We used metagenomics to investigate the biodiversity, functional variability, and genetic exchange among the endosymbionts of 12 lucinid host species from across the globe. We report a cosmopolitan symbiont species, Candidatus Thiodiazotropha taylori, associated with multiple lucinid host species. Ca. T. taylori has achieved more success at dispersal and establishing symbioses with lucinids than any other symbiont described thus far. This discovery challenges our understanding of symbiont dispersal and location-specific colonization and suggests both symbiont and host flexibility underpin the ecological and evolutionary success of the lucinid symbiosis.
Acknowledgements (from the paper)
We thank Gustav Paulay (FLMNH, Gainesville, FL), Amanda Bemis, and John Taylor (NHM, London) for providing preserved samples. John Taylor also provided high-resolution images of lucinid clams for Fig. 2. We thank Liz Hambleton (University of Vienna), Miriam Weber, Christian Lott, Boris Unger, Dorothée Makarow, Natalie Elisabeth (University of California, Berkeley), Diana Chin, Tim Oortwijn, and Arthur Curry for providing fresh samples. We also thank Thalassa for 300 million y of sheltering lucinids. We are grateful to Stephanie Markert for providing access to the proteomics data, Ipek Yamin Meric and Nora Grossschmidt for helping with DNA extractions, Marina DeLeón for help with fieldwork and travels in Costa Rica, Cassandra Ettinger for early discussions of the study, and Guillaume Jospin for support with the bioinformatic analyses. We thank Xavier Didelot for discussions on the recombination analysis. L.G.E.W. was supported by Grant GMBF5603 from the Gordon and Betty Moore Foundation (J.A.E. was Principal Investigator on the same grant). The work was also supported by the European Research Council Starting Grant EvoLucin and a Vienna Research Grant for Young Investigators from the Vienna Science and Technology Fund (WWTF, VRG14–021) and the Austrian Academy of Sciences. Sequencing was carried out at the DNA Technologies and Expression Analysis Cores at the University of California, Davis Genome Center (supported by NIH Shared Instrumentation Grant 1S10OD010786-01), the Biomedical Sequencing Facility at CeMM (https://www.biomedical-sequencing.org/) in Vienna, Austria, and at the Joint Microbiome Facility (JMF) of the Medical University of Vienna and the University of Vienna (project ID 1911-1). Thanks to Petra Pjevac and Gudrun Kohl for sample processing at JMF. The gill embedding was performed by the Histopathology Facility at Vienna BioCenter Core Facilities, a member of the Vienna BioCenter, Austria.
Funding: Gordon and Betty Moore Foundation (for the part from my lab at least). Part of a collaboration with STRI.