My son goes to DCCNS preschool. They had a fundraiser yesterday at Capay Organic Farm in Capay (this is the place that the Farm Fresh to You folks have their farm). It was a nice time. Party started at 2 pm, they had tractor rides, kite flying, ladybug releasing, strawberry picking, and more. And then there was a raffle (the fundraiser part) and tomato planting to take home and a piñata. The farm was beautiful, the people there were great, and I am glad we get food from there (we are subscribers to their CSA). It was good for all the kids to see where some food comes from … Here are some pics. https://picasaweb.google.com/s/c/bin/slideshow.swf
Author: Jonathan Eisen
Dueling microbial diversity talks at #UCDavis on May 2 #symbioses #microbiology
Here is a storification of the dueling microbial diversity talks that happened at UC Davis on Wednesday May 2.
http://storify.com/phylogenomics/dueling-microbial-diversity-talks-at-ucdavis.js?template=slideshow[View the story “Dueling microbial diversity talks at #UCDavis” on Storify]
How complete do microbial genomes have to be for metabolic predictions (to be useful)?
OK. Got a question for the blogo-twitto-webosphere.
In this day an age of rapid shotgun sequencing of genomes, many people are moving away from finishing the genomes. As some may know, I was a co-author many years ago on a paper arguing for the need to finish (rather than just shotgun sequence) microbial genomes for many scientific questions. But as sequencing costs continued to plummet, the relative cost of shotgun sequencing genomes kept going down while the cost of finishing genomes did not change much. So. two years ago I posted to my blog a question regarding this: “Wanted:Feedback on Importance of Finishing (Microbial) Genomes” and got a lot of useful feedback. And eventually, I came around to the argument that finishing was unnecessary for many purposes (and even got some props for admitting I was wrong).
Well, I am back. Now I am arguing to some colleagues that if we want to make metabolic pathway predictions and metabolic models for genomes, we probably don’t need finished genomes. But alas, I have no evidence to back that up. And in fact, I am not really sure anyway. So I am asking everyone and anyone out there … does anyone have data/evidence/opinions about whether there will be much difference in metabolic predictions one would make for an organism based upon a complete genome vs. a shotgun assembly of a genome generated by Illumina sequencing?
Thanks
Storification of some comments:
http://storify.com/phylogenomics/completeness-of-microbial-genomes-for-metabolic-pr.js[View the story “Completeness of microbial genomes for metabolic predictions” on Storify]
Draft of a Proposal for a UC #OpenAccess policy – comments wanted
Just got sent this email
Dear Colleagues,
On behalf of the Academic Senate Library Committee (ASLC), I am asking for your comments on the attached proposed Open Access Publishing Policy for the University of California.. All faculty, including Academic Federation members are invited to post their comments on the Academic Senate web-forum site at http://academicsenate.ucdavis.edu/Forums/index.cfm?Forum_ID=67. Please go to this site to submit your feedback.
Briefly, the issue is this: the faculty of the University of California, in conjunction with the University Committee on Libraries and Scholarly Communication (UCOLASC), is proposing a new OPEN ACCESS PUBLISHING POLICY that will apply to the dissemination of all scholarly work. UCOLASC is seeking feedback from all campuses on this issue in order to inform a final version of the policy which will be presented to the Universitywide Academic Senate sometime this calendar year.
The ASLC would appreciate your comments by Wednesday, May 9, 2012. Your ideas will then be shared with UCOLASC in time for its May 25th meeting. The web-forum will remain open substantially past May 9, and we will endeavor to include as many comments up to May 25 as possible.
Sincerely,
Brian H. Kolner
Academic Senate Library Committee
The relate to a draft of a proposal for a new Open Access Publishing Policy being circulated at the University of California. The draft of the proposal can be found here.
UC Davis (and I presume other UCs) are now soliciting comments on the proposal. I would love to here / read comments from anyone. Personally, I think the policy is way to weak as it allows exceptions to be granted …
SEEDS: BioBlitz information #DavisCA #Nature #Biodiversity
DAVIS BioBlitz!
“In Honor of Mother Earth”
May 13, 2012
(Mother’s day)
Putah Creek Reserve
Discover the Many Plants and Animals in Putah Creek Reserve, and learn about ways to preserve and protect the earth!
Free Activities for All Ages!
Hosted by the UC Davis SEEDS Chapter,
a program of the Ecological Society of America
What is a BioBlitz?
A BioBlitz consists of activities to draw attention to the biodiversity of plants, and animals that live in Putah Creek Reserve. These activities include:
Species Counts:As a group we will count and document as many species as we can in Putah Creek: plants, birds, mammals, reptiles, amphibians, insects, and spiders.
Activities for the public: Kids and adults get to observe, explore, discover, touch, ask questions about, and enjoy the flora and fauna of Putah Creek Reserve.
Species found during the BioBlitz will be documented using iNaturalist and recorded in a National database. (For more information about iNaturalist visit iNaturalist.org).
**Volunteers needed for the day of the event?***
Are you an expert at identifying plant, fungi, or animal species?
Interested in gaining more experience with identification?
Just want to help out and learn something new about ecosystems near you?
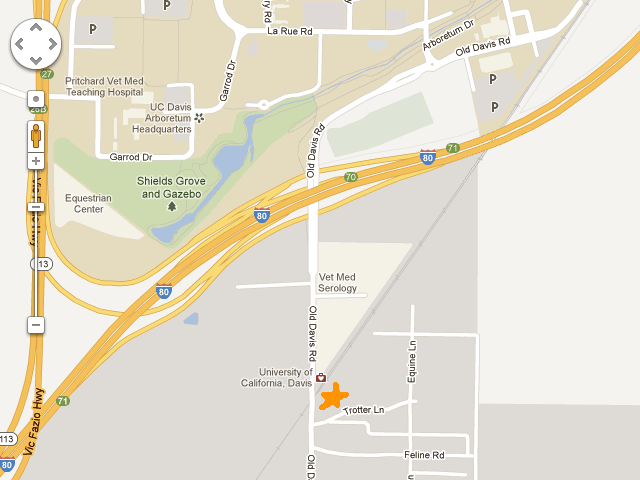
Location: Putah Creek Reserve,
BioBlitz headquarters will be set up on Old Davis Road- Look for BioBlitz Signs!
Social Networks and Scientists: Chronicle for Higher Education Article
Quick post here.
There is a new article in the Chronicle for Higher Education in which I am quoted: Social Networks for Academics Proliferate, Despite Some Scholars’ Doubts
The article discusses many connected topics relating to the use of social media by scientists – though it does not make clear how everything is connected perhaps. Anyway the author talked to me about Mendeley and various uses of Mendeley and I told her about an effort to create a Mendeley collection of my father’s papers. The article also discussed LinkedIn, Academia.Edu, Twitter and other social media systems.
Some quotes
Jonathan A. Eisen, a professor of medical microbiology and immunology at the University of California at Davis, used Mendeley to distribute the research papers that his father, Howard J. Eisen, a researcher at the National Institutes of Health, published before he died, in 1987. After struggling to free papers locked behind pay walls, Jonathan Eisen compiled the articles and posted nearly all of them on a Mendeley page he had created for his father.
Mr. Eisen, a self-described “obsessed open-access advocate,” described the impact in a blog post last year: “Thanks to the social features of Mendeley, more and more people will see and have access to those papers, thus ensuring that they do not wallow in never-never land but continue to have some potential impact on science and society.”
Perhaps most important from my point of view – I love the picture of me taken by Max Whittaker.
Summary of responses to question about metrics for density in phylogenetic trees
I posted a question to Twitter and Facebook about metrics for assessing density in a phylogenetic tree. Here is a “Storification” of the responses. Thanks for the help all.
Any other suggestions welcome in comments … http://storify.com/phylogenomics/metrics-to-quantify-density-of-taxa-sampling-in-a.js?template=slideshow[View the story “Metrics to quantify density of taxa/sampling in a phylogenetic tree” on Storify]
Conference: LosAngeles 5th International Conference of Anaerobic Protists Sep6-9
From EVOLDIR
Dear Colleagues,
Please visit the website for the 5th International Conference of Anaerobic
Protists – https://sites.google.com/site/2012icap/home – which will be
in Los Angeles, California Sept 6-9, 2012 for information about this
important and exciting conference.You will be able to register and submit abstracts (for both talks
and poster presentations) between now and July 1. Detailed information
about how to do this can be found on the website. Additional information
regarding the goals and structure of the meeting, travel and housing,
etc. is also available on the website.Your participation in this meeting is important to make it a success! We
hope to see you in LA in September!Best regards,
Patricia Johnson (on behalf of the organizing committee)
Professor, UCLA
Seminar 5/7 at #UCDavis Reed Cartwright “Evolutionary models of mutation & variation for genomic data”
Genetics Seminar
“Evolutionary models of mutation and variation for genomic data”
Speaker: Reed Cartwright
Arizona State University | Biodesign Institute
Monday, May 7, 2012
4:10 PM
1022 Life Sciences
UC Davis/Berkeley Joint Colloquium, 5/3. Rasmus Nielsen “Statistical Problems in Analysis of Next-Gen Sequencing Data”
The 2012 UC BERKELEY / UC DAVIS JOINT STATISTICS COLLOQUIUM
Please join us for the 2012 Berkeley / Davis Joint Colloquium. This year’s Colloquium will feature a talk at 4:10pm by Prof. Rasmus Neilsen, followed by a reception in the statistics lounge. Grad students are also invited to the Berkeley / Davis Grad Student Colloquium from 5:30-6:30pm.
Coffee: 3:30pm, Statistics lounge (MSB 4110, 4th floor)
Seminar: 4:10 pm, Colloquium Room (MSB 1147)
Reception: 5:30 pm, Statistics lounge (MSB 4110, 4th floor)
Speaker: Prof. Rasmus Neilsen
Dept of Integrative Biology & Statistics, UC Berkeley
Title: Statistical Problems in the Analysis of Next-Generation Sequencing Data
Abstract: The biological sciences have been transformed by the emergence of Next-Generation Sequencing (NGS) technologies providing cheap and reliable large scale DNA sequencing. These data allow us to address biological research questions that previously were considered intractable, but also raise a number of new statistical and computational challenges. The data contain errors that need careful attention, and the appropriate likelihood functions are usually not computationally accessible, because of the size of the data sets. I will discuss some solutions to these problems and illustrate them in the analyses of several different data sets. In one study we sequenced all protein coding genes of 2000 individuals to identify mutations associated with Type 2 Diabetes. In a second project we used similar sequencing techniques to identify the genetic causes of altitude adaptation in Tibetans. In the third study I will discuss, we sequenced the first Aboriginal Australian genome to elucidate the history and origins of Aboriginal Australians.
Graduate Student Colloquium : Please see the attached flier for details of the Graduate Student Colloquium, which will feature talks by Andrew Farris (UC Davis) and Vincent Yates (UC Berkeley).