Just got alerted to this new (open access) paper that seems like it should be of interest to those working on plant microbiomes Scientia Agricola – Exploring interactions of plant microbiomes. It has some useful summaries of work that has been done on plant microbiomes.
Eisen Lab Blog
Do preprints count for anything? Not according to Elife & G3 & some authors ..
Well, just got pointed to this paper: Metagenomic chromosome conformation capture (meta3C) unveils the diversity of chromosome organization in microorganisms | eLife by Martial Marbouty, Axel Cournac, Jean-François Flot, Hervé Marie-Nelly, Julien Mozziconacci, Romain Koszul. Seems potentially really interesting.
It is similar in concept and in many aspects to a paper we published in PeerJ earlier in the year (see Beitel et al., 2014 Beitel CW, Froenicke L, Lang JM, Korf IF, Michelmore RW, Eisen JA, Darling AE. (2014) Strain- and plasmid-level deconvolution of a synthetic metagenome by sequencing proximity ligation products. PeerJ 2:e415 http://dx.doi.org/10.7717/peerj.415.
Yet despite the similarities to our paper and to another paper that was formally published around the time of ours, this new paper does not mention these other pieces of work any where in the introduction as having any type of “prior work” relevance. Instead, they wait until late in their discussion:
Taking advantage of chromatin conformation capture data to address genomic questions is a dynamic field: while this paper was under review, two studies were released that also aimed at exploiting the physical contacts between DNA molecules to deconvolve genomes from controlled mixes of microorganisms (Beitel et al., 2014; Burton et al., 2014).
Clearly, what they are trying to do here is to claim that since they paper was submitted before these other two (including ours) was published, that they should get some sort of “priority” for their work. Let’s look at that in more detail. Their paper was received May 9, 2014. Our paper was published online May 27 and the other related paper by Burton et al. was published online May 22. In general, if a paper on what your paper is about comes out just after you submit your paper, while your paper is still in review, the common, normal thing to be asked to do is to rewrite your paper to deal with the fact that you were, in essence, scooped. But that does not really appear to be the case here. They are treating this in a way as “oh look, some new papers came out at the last minute and we have commented on them.” The last minute would be in this case, 6 months before this new paper was accepted. Seems like a long time to treat this as “ooh – a new paper came up that we will add a few comments about”.
But – one could quibble about the ethics and policies of dealing with papers that were published after one submitted one’s own paper. From my experience, I have always had to do major rewrites to deal with such papers. But maybe E-Life has different policies. Who knows. But that is where things get really annoying here. This is because it was May 27 when our FINAL paper came out online at PeerJ. However, the preprint of the paper was published on February 27, more than two months before their paper was even submitted. So does this mean that the authors of this new paper do not believe that preprints exist? It is pretty clear on the web site for our paper that there is a preprint that was published earlier. Given what they were working on – something directly related to what our preprint/paper was about, one would assume they would have seen it with a few simple Google searches. Or a reviewer might have pointed them to it. Maybe not. I do not know. But either way, our preprint was published long before their paper was submitted and therefore I believe they should have discussed it in more detail.
Is this a sign that some people believe preprints are nothing more than rumors? I hope not. Preprints are a great way to share research prior to the delays that can happen in peer review. And in my opinion, preprints should count as prior research and be cited as such. I note – the Burton group in their paper in G3 also did not reference our preprint in what I consider to be a reasonable manner. They add some comments in their acknowledgements
While this manuscript was in preparation, a preprint describing a related method appeared in PeerJ PrePrints (Beitel et al. 2014a). Note added in proof: this preprint was subsequently published (Beitel et al. 2014b).
Given that our preprint was published before their paper was submitted too, I believe that they also should have made more reference to it in their paper. But again, I can only guess that both the Burton and the Marbouty group just do not see preprints as being respectable scientific objects. That is a bad precedent to set and I think the wrong one too. And it is a shame. A preprint is a publication. No – it is not peer reviewed. But that does not mean it should not be considered part of the scientific literature in some way. I note – this new paper from the Marbouty group seems really interesting. Not sure I want to dig into it any deeper if they are going to play games with the timing of submission vs. published “papers” as part of how they are positioning themselves to be viewed as doing something novel.
Buzzfeed includes microbiomes in "77 facts" but alas, gets the facts (and the math) wrong
Well, I admit it – I clicked on this Buzzfeed link someone posted on Facebook – 77 Facts That Sound Like Huge Lies But Are Actually Completely True. There are some pretty funny and interesting things on the list. However, I note – I did not click on the link per se for entertainment. I clicked on it to see if there was anything about microbes on the list. And happily, there was. Alas, what there was, was, well, a bit wrong:
Let’s start with #67. “There is 10 times more bacteria in your body than actual body cells“. Well, alas, this is a nice bit of information. But it is not based on facts. See Peter Andrey Smiths wonderful article in the Boston Globe about this issue. The Buzzfeed article links to a 2010 Discover Magazine bit about this topic. So they were trying to use facts. But alas, that was not based on facts itself.
Then, let’s go to #68. “And 90% of the cells that make us up of aren’t human but mostly fungi and bacteria.“. So – I am at a bit of a loss on this. First, isn’t this really just rehashing #67 with the addition of fungi to the story? Also – I note the math here is weird. #67 would lead one to conclude that 90.9% of the cells in your body are bacterial (10:1 = 10/11 = 90.9%). So then if one adds fungi to the picture the percent of cells in our bodies that are not human goes DOWN to 90%. How does that work exactly?
So not only do they get the 10x as many cells fact wrong, they do something really weird by basically repeating #67 and then doing some weird math.
Oh well, glad they have something on microbiomes. Too bad it came out a bit wrong.
postdoctoral position openings in microbial evolution and ecology
Dear colleagues,
Below and attached is information about two new postdoctoral positions in my lab. Please help spreading the word to potential candidates.
Thank you,
Ramunas Stepanauskas
Two postdoctoral research scientist positions in microbial evolution and ecology
Summary
Two postdoctoral positions are available in Dr. Stepanauskas’ laboratory. One position is focused on the deep evolution of Bacteria and Archaea and the genome content of the “microbial dark matter”. Another position is focused on chemolithoautotrophy in the deep ocean and hydrothermal systems. The hired scientists will be engaged in large, collaborative projects, and will employ single cell genomics and other advanced molecular biology and biogeochemistry research tools. Anticipated employment duration: 2 years, with potential extension. Bigelow Laboratory’s new campus is located in scenic, coastal Maine with abundant opportunities for outdoor and cultural activities. It is about an hour drive from Portland and a 3-hour drive from Boston.
Responsibilities
Hired scientists will be responsible to lead computational analyses of large single cell genomics, community “omics” and biogeochemical data sets, prepare manuscripts, supervise undergraduate students, and assume gradually increasing responsibilities in project management. These positions also include opportunities for field expeditions and laboratory analytical work.
Requirements
Candidates must have a PhD degree in microbiology, evolutionary sciences, bioinformatics, computational biology, or a related field. Prior experience in computational analyses of large microbial genomic data sets will be necessary for this opportunity. Candidates interested in these positions must be highly motivated, willing to learn and demonstrate initiative in assigned tasks. Excellent written and verbal communication skills and willingness to work in teams composed of field, laboratory and computational scientists are crucial.
Research overview: Deep evolution of Bacteria and Archaea
Hired scientist will be engaged in a major effort to improve our understanding of the genealogy of the Bacteria and Archaea by a large-scale genomic analysis of those deep evolutionary branches (phyla) that contain no cultivated representatives. Popularly referred to as Microbial Dark Matter (MDM), these enigmatic limbs near the base of the tree of life constitute a large fraction of biological diversity on our planet; yet, we learned of their existence only in 1990’s. Current knowledge about MDM is mostly limited to sequences of its small subunit ribosomal RNA genes, primarily due to former technology limitations. Recent advances in single cell genomics technology, however, have enabled us to bridge this knowledge gap – one that is essential to the genealogy of life – by providing access to the complete genomic blueprints of microorganisms without the need for cultivation. The following, general hypotheses will be tested: 1) The extant number of major (phylum-level) evolutionary branches of Bacteria and Archaea is significantly greater than the current consensus; 2) Extant cellular life forms three distinct domains: Bacteria, Archaea and Eukaryotes; 3) The early evolution of Bacteria and Archaea followed a progression of bifurcating divergences rather than a single radiation from the last universal common ancestor; 4) Inter-domain and inter-phylum horizontal gene transfer (HGT) had a significant impact on the early evolution of Bacteria and Archaea; 5) Evolutionary divergence in the early history of Bacteria and Archaea co-occurred with the colonization of and adaptations to novel environments; 5) The habitable subsurface is a repository for early evolutionary history of Bacteria and Archaea. Primary field study sites include the Kaapvaall Craton of South Africa; the Sanford Underground Research Facility in South Dakota; and the Death Valley Regional Flow System in Nevada.
Research overview: Chemolithoautotrophy in the dark ocean
An increasing body of evidence suggests the significance of chemolithoautotrophy in the dark ocean water column and hydrothermal systems. However, the specific energy sources, metabolic pathways and microbial taxonomic groups remain poorly understood. The hired scientist will assume a leading role in a) A global inventory of chemoautotrophs in the dark ocean water column; and b) An integrated study of energy metabolism, carbon fixation, and colonization mechanisms in chemosynthetic microbial communities at deep-sea vents. The following general hypotheses will be tested: 1) Multiple prokaryote taxonomic groups found in the dark ocean contain chemoautotrophic metabolic pathways; 2) Both known and previously unrecognized chemoautotrophy pathways are present in dark ocean’s prokaryotes; 3) Dark ocean chemoautotrophs are broadly distributed around the globe, with biogeographic patterns determined by the isopycnal movement of water masses, water mass age, and the downward flux of organic matter; 4) Diverse chemoautotrophy pathways are expressed in the dark ocean. During the course of the project, single amplified genomes (SAGs) will be generated from several diffuse-flow hydrothermal systems and from all major intermediate and deep water masses around the globe, representing most major taxonomic groups of bacteria and archaea that are known to be present in the dark ocean. Whole genome sequencing will be performed on a subset of SAGs, enabling detailed annotation of chemoautotrophy pathways. Metagenomic and metatranscriptomic fragment recruitment will be used to determine global patterns of chemoautotroph distribution and chemoautotrophy pathway expression.
How To Apply
Please visit www.bigelow.org for more information and method of application, position reference # PD-2015-1 for the Ecology position and PD-2015-2 for the Evolution position. Application deadline: February 1, 2015.
—
Ramunas Stepanauskas, Ph.D.
Senior Research Scientist
Director of the Bigelow Laboratory Single Cell Genomics Center
Crosspost – papers for sale
Crossposting from ICIS Blog (even though this is short – always good to reveal where things are posted first).
Well, this is much more elaborate than I could ever have imagined: For Sale: “Your Name Here” in a Prestigious Science Journal – Scientific American. Seems that there are services out there to help people write, in essence, bogus scientific papers filled with pithy somewhat reasonable sounding phrases about certain topics. Seems we could all use some more comprehensive full text analyses of papers to try and flag such activities.
Quick post – papers for sale
Well, this is much more elaborate than I could ever have imagined: For Sale: “Your Name Here” in a Prestigious Science Journal – Scientific American. Seems that there are services out there to help people write, in essence, bogus scientific papers filled with pithy somewhat reasonable sounding phrases about certain topics. Seems we could all use some more comprehensive full text analyses of papers to try and flag such activities.
Wanted – info on workshops in and around Northern California on bioinformatics & genomics
I have been asked by multiple students about this topic and figured I would just put it out there. I am trying to compile information on workshops and short courses on topics relating to genomics or bioinformatics that are in the general area of Northern California (you know, Davis, Sacramento, SF, Berkeley, Stanford, Santa Cruz, etc). This would include using specific tools (e.g., R, Galaxy, QIIME, and more) and specific fields (e.g., ecology, microbiology, genetics, plant biology). Any information would be appreciated.
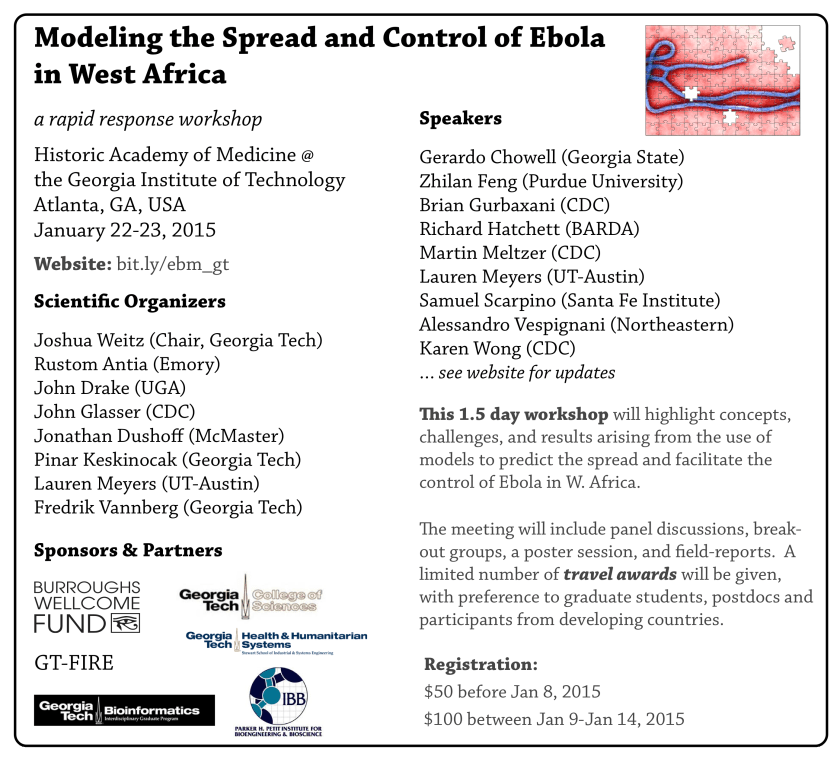
Meeting announcement: “Modeling the Spread and Control of Ebola in W. Africa”, Atlanta, GA, Jan 22-23, 2015
Of possible interest:
Modeling the Spread and Control of Ebola in W. Africa
Historic Academy of Medicine @ Georgia Tech
Atlanta, GA, USA
Jan 22-23, 2015
Website: bit.ly/ebm_gt
Please see attached for an image and pdf meeting flyer. The organizing committee includes Pinar Keskinocak and Fred Vannberg from Georgia Tech, as well as Rustom Antia (Emory), John Drake (UGA), Jonathan Dushoff (McMaster), John Glasser (CDC) and Lauren Meyers (UT-Austin).
The meeting will begin at 8:30am on Jan 22 and conclude by 1:30pm on Jan 23. Note that there is a $50 registration fee for all participants before Jan 8, 2015. This fee includes access to all events, as well as breakfasts, lunches and coffee breaks for the 1.5 day meeting.
We have a maximum capacity of 200 individuals on-site. We also expect to be able to offer travel awards targeted to young researchers and participants from developing countries – preference will be given to those who intend to present a poster at the meeting.
Please distribute to colleagues, questions should be addressed to:
ebola-modeling-workshop
Great idea from Nicole King – Lists of Women Speakers – more examples wanted
Just been pointed to this compilation from Nicole King (a brilliant professor at Berkeley): Lists of Women Speakers | The King Lab.
She has put together a collection of a few links:
As a resource for those who are interested in increasing the number of female speakers at their scientific conferences, seminar series, etc., I offer here a compendium of female speakers from diverse fields. I’ve started with lists that I could find easily, but if you know of additional lists, please feel free to send along the links. I would also be happy to maintain similar lists for underrepresented minorities if any such lists exist.
These are the lists she maintained:
- University of Minnesota, Distinguished Scientists and Engineers from Underrepresented Groups
- Women scientists that give awesome seminars
- Celebrating Women in Science Symposium
- Indiana University Joan Wood Lecture Series
- American Society of Cell Biology WICB Speaker Referral Service
- Neuroscience: Annie’s List
- Theoretical/computational chemistry, material science, and biochemistry
- Synthetic Biology
- Prominent ecologists
- Geoscientists
- American Physical Society Women Speakers List
Some responses from Twitter
@duffy_ma @phylogenomics a few months ago I (& many others) sent in some suggestions to @kejames who I think was compiling a similar list.
— Opposing Thumb (@OpposingThumb) December 18, 2014
//platform.twitter.com/widgets.js
@jkpfeiff @phylogenomics Are female plant virologists included? I know two and they aren’t on the list… Hélène Sanfaçon and D’Ann Rochon.
— David Joly (@idjoly) December 18, 2014
//platform.twitter.com/widgets.js
@idjoly @phylogenomics Yes, we want plant virologists for sure! I just added them. Thanks!
— Julie Pfeiffer (@jkpfeiff) December 18, 2014
//platform.twitter.com/widgets.js
@phylogenomics Came across @EMBOcomm Women in Life Sciences database the other day. Covers 17 disciplines: http://t.co/Ul7sshqxdj @duffy_ma
— Rachel Brenchley (@rachelcwalton) December 18, 2014
//platform.twitter.com/widgets.js
@OpposingThumb @duffy_ma @phylogenomics Here is my working list of top women of color in biology: http://t.co/QsvMpGKVvM
— Karen James (@kejames) December 18, 2014
//platform.twitter.com/widgets.js
@phylogenomics We’re working on a project in this area too https://t.co/uHmfBMLkLi
— Innovation Women (@WomenInno) December 19, 2014
//platform.twitter.com/widgets.js
@OpposingThumb @duffy_ma @phylogenomics Another 1, @kejames Diana Bautista frm @Cal, fresh off #SfN Young Invest awrd http://t.co/gU5hW8P6C4
— Sam Diaz-Munoz (@evolcoop) December 19, 2014
Call for Participant Applications: Sustaining Biological Infrastructure
Of possible interest …
 |
| Dear NSF Division of Biological Infrastructure Principal Investigator:
The Ecological Society of America (ESA) announces the training course “Sustaining Biological Infrastructure (SBI): Strategies for Success – A Short Course for Project Directors.” Please see the call for applications below for more details about the course and how to apply. We hope that this course will be of particular interest to you and your colleagues. If you have any questions please contact us! Sincerely, |