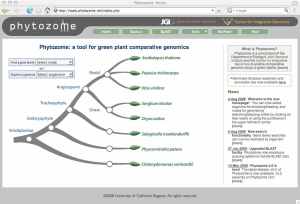
 I spent the last few days at a “retreat” for the Joint Genome Institute and heard about a few things there worth sharing with everyone. I will try and post about some of them in the next few days. Here is one. The JGI and the Center for Integrative Genomics have made a pretty cool tool for comparative analyses of plant genomes. It is called Phytozome and has a variety of simple and nice features. JGI is doing more and more work on plant genomes as part of their energy research and I think Phytozome could turn into a good place to go to get the latest plant genome information. Go to http://www.phytozome.net to see the real thing.
I spent the last few days at a “retreat” for the Joint Genome Institute and heard about a few things there worth sharing with everyone. I will try and post about some of them in the next few days. Here is one. The JGI and the Center for Integrative Genomics have made a pretty cool tool for comparative analyses of plant genomes. It is called Phytozome and has a variety of simple and nice features. JGI is doing more and more work on plant genomes as part of their energy research and I think Phytozome could turn into a good place to go to get the latest plant genome information. Go to http://www.phytozome.net to see the real thing.
Skip to content


That’s a really nice resource! Great interface, too. Looking fwd to reading the paper describing the clustering algorithm. Is it distance-based? A similar tool is < HREF="http://nypg.bio.nyu.edu/orthologid" REL="nofollow">OrthologID<> that uses a character-based phylogenetic framework to establish orthology. Chiu et al. (2006) OrthologID: automation of genome-scale ortholog identification within a parsimony framework. <>Bioinformatics<> 22(6):699-707. [< HREF="http://dx.doi.org/10.1093/bioinformatics/btk040" REL="nofollow">free full text<>]
LikeLike
dont know any more — just posted the information about it — I am not involved in making it
LikeLike
Wow, clustering of extant genes to get ancient genome data, and all in one site! I’m sure this will be a great resource for my work. I also love the data retrieval service BioMart at Phytozome.
LikeLike