Just because you have data that could be plugged into a phylogenetic analysis does not mean it makes sense to do so. Case in point – the following paper:
A Differentiation-Based Phylogeny of Cancer Subtypes by Riester M, Stephan-Otto Attolini C, Downey RJ, Singer S, Michor F.
In this paper the authors take gene expression data from various cancer samples/cell lines and then they build phylogenetic trees from the data. See example below:
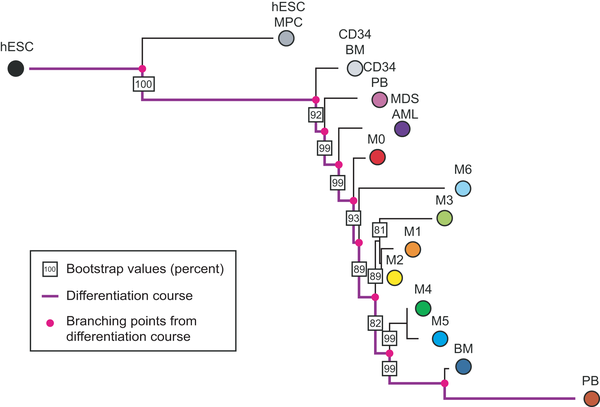
Figure 2. A phylogeny of acute myeloid leukemia (AML) subtypes. According to the French-American-British (FAB) classification, AML samples are classified into seven different types according to their level of differentiation (see Table 1). Expression data from 362 AML patients and 7 Myelodysplastic Syndrome (MDS-AML) patients is used to construct a phylogeny of these leukemias. We include expression data of human embryonic stem cells (hESCs), CD34+ cells from bone marrow (CD34 BM) and peripheral blood (CD34 PB), and mononuclear cells from bone marrow (BM) and peripheral blood (PB). The differentiation pathway from hESCs to mononuclear cells from peripheral blood is represented in purple, and the common ancestors of subtypes are shown as pink dots. The bootstrap values of branches are indicated by boxed numbers, representing the percentage of bootstrapping trees containing this branch. The ranking of AML subtypes identified by the phylogenetic algorithm corresponds with the differentiation status indicated by the FAB classification. The M6 subtype, represented by only 10 samples in our dataset, has the least stable branch, leading to lower bootstrap values for those branches where it can alternatively be located.
The pictures are pretty. They make some sense biologically. The paper has some very interesting parts and I do not want to suggest that the paper is not useful. But it makes no sense to me to use a phylogenetic approach to analyze this data. Phylogenetic methods are about reconstructing history of evolutionary lineages. They should not be doing that here as far as I can tell since the cancers are from different people with different histories and what they make be looking at is convergent / developmental similarities in the cancer samples. But they are not looking at history per se. And thus it is not appropriate to use algorithms that use phylogenetic methods:
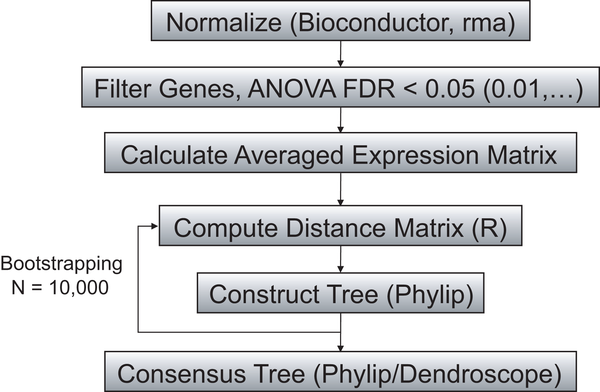

It just makes no sense to me to use a phylogenetic method instead of some sort of clustering method in the step where it says “construct tree” in their flow diagram. Sure phylogenetic methods can make nice pictures. But they should only be used when the underlying data has a history that is reflected in the model/assumptions of the phylogenetic method. I could, for example build a phylogeny of cities based on various metrics. But would that make sense? Most likely not. Don’t get confused by the fact that similar things group together in the same part of a phylogenetic tree to thinking that that means that a phylogenetic model is right for your data.
I may be obsessed with phylogeny but that obsession applies to applying phylogenetic methods to data with histories that are approximated by the methods being used … and this paper seems to not be doing that …
Hat tip to Eric Lowe, an undergrad in my lab for showing me this paper.
I note – this does not mean that phylogenetic methods cannot be applied to cancer studies. Case in point – this paper:
Estimation of rearrangement phylogeny for cancer genomes by Greenman CD, Pleasance ED, Newman S, Yang F, Fu B, Nik-Zainal S, Jones D, Lau KW, Carter N, Edwards PA, Futreal PA, Stratton MR, Campbell PJ.
In this paper the authors focus on mutations in cancer cells and they use phylogenetic methods to determine the order in which genomic changes happen in these cancer cells. This seems to be an excellent use of phylogenetic / phylogenomic methods.
So – lesson of the day – phylogenetic methods should be used on data with a phylogenetic history. Not so complicated. But pretty important.








