Another request to Twitter and some very useful responses and discussion. I have compiled the responses via Storify.
Author: Jonathan Eisen
Request for suggestions for bacterial genome annotation tools (Storify of some answers)
Posted a request to Twitter about tools for genome annotation and got a collection of useful responses. I have compiled them here via Storify.
#Badomics word of the year already? Yup. The ‘Consciousome’ from Deepak Chopra
Oh my. As many out there know I have a thing with “badomics” words. These are words someone has invented where they have added the ome suffix on to the word to try and capture some of the hype of genomics. And though many many people do this, the ones I call “badomics” words are the ones where the ome addition is all hype and no use. And this AM I was informed of a doozy via Twitter:
@phylogenomics Have you seen ‘consciousome’? http://t.co/oUJf8Sumyy
— Blanca Himes (@blancahimes) January 17, 2014
And so I went to the link and found this: Self-Directed Biological Transformation Initiative — A New Frontier ‘Consciousome’ | Deepak Chopra and Rudolf E. Tanzi. And what I found there was almost a textbook example of how to create a badomics word. Here is the critical paragraph
The current frontier in brain research, to map the entire connectivity of the brain, the so-called connectome project, points to an even more exciting frontier, the “consciousome.” This takes the brain to the next level, where we need to explore how our conscious choices may liberate us from biology-as-destiny. Our conscious choices and reactions to life experiences remodel the neural circuitry of our brains and, now, we need to explore the effects on our genomes.
Just what is the consciousome? I cannot tell. But it has something to do with genomics. And with consciousness. Hmm. They try to clarify later on
Where the connectome is like a diagram of all the telephone wiring in a city, the consciousome embraces the conversations taking place using the wiring.
Not really helping. So – what does it have to do with genomics? Unclear until later
By opening up the genetic doorway to consciousness, however, we take a leap forward. For example, recent studies indicate that meditation can have a strong effect on the length of chromosome telomeres, the nucleotide sequences that protect chromosomes from the deterioration linked to aging. That these beneficial effects occur immediately indicates just how responsive genetic activity is to mind-body interventions — something never previously suspected.
So – the consciousome is the affect of conscious thought on the genome? Whether or not you think this is something worth studying, the slapping on of the ome term on the end of consciousness is a self directed thought that should never have been done. There is no entity that is the consciousome. The term makes no sense. It is certainly a badomics word of grand magnificence.
I note – there is some seriously bizarre other stuff in this article like the following:
In our Self-Directed Biological Transformation Initiative, a 100-subject study to map biological transformation, leaders in the fields of healthcare, art, business, environment, sports, entertainment, science and technology will be given a protocol to measure new genetic activity, searching for positive evolutionary effects
What? How exactly are they searching for “positive evolutionary effects” and what does that even mean? And why do this with the “leaders in the fields …”. How are they supposed to be different?
And also
This “soft inheritance,” in which the parents’ life experiences and behavior may directly influence the genome of their offspring (transmitted via the epigenome), is arguably the most profound “living legacy” we can pass along to our children.
So now they have gone from a few studies in mice showing epigenetic inheritance of a few things to the “most profound living legacy we can pass along to our children”. Uggh. Talk about overselling genomics – this is overselling a word / field that does not even exist yet and the word certainly should not exist ever. Personally, I think brain science is very very interesting and important. Let’s not cloud it up by coining new fuzzy terms, trying to capture the hype of genomics, and overselling the science. And for this, Deepak Chopra, Rudolf Tanzi and their coauthors Tara Sheahan, Gina Murdock, and
Glenda Greenwald are winners of the coveted “Worst New Omics word award.”
At #UCDavis today: Erin Mordecai “Unexpected responses of disease to global change”
DEPARTMENT OF EVOLUTION AND ECOLOGY
RECRUITMENT SEMINAR
ECOLOGIST
Dr. Erin Mordecai
NSF Postdoctoral Research Fellow
University of North Carolina and
North Carolina State University
“Unexpected responses of disease to global change"
Thursday, January 16th, 2014
2:10pm
1022 Life Sciences Building
Faculty Host: Professor John Stachowicz, Department of Evolution and Ecology
jjstachowicz
Carin Bondar video on "Organisms do Evolve" based on Miley Cyrus’ Wrecking Ball
I love Carin Bondar.
A little bit about PhyloSift: phylogenetic analysis of genomes and metagenomes
New paper from people in the Eisen lab: PhyloSift: phylogenetic analysis of genomes and metagenomes [PeerJ].
Basically, the concept behind Phylosift is to provide for high quality, automated, high throughput phylogeny-driven analysis of metagenomic sequence data. The software was developed openly on github and has been available in some form for more than a year. Aaron, Holly, Erick and I have discussed it extensively in various talks around the world and thus we assume some are already familiar with it.
This project was coordinated by Aaron Darling, who was a Project Scientist in my lab and is now a Professor at the University of Technology Sydney. Also involved were Holly Bik (post doc in the lab), Guillaume Jospin (Bioinformatics Engineer in the lab), Eric Lowe (was a UC Davis undergrad working in the lab) and Erick Matsen (from the FHCRC).
Abstract:
Like all organisms on the planet, environmental microbes are subject to the forces of molecular evolution. Metagenomic sequencing provides a means to access the DNA sequence of uncultured microbes. By combining DNA sequencing of microbial communities with evolutionary modeling and phylogenetic analysis we might obtain new insights into microbiology and also provide a basis for practical tools such as forensic pathogen detection.
In this work we present an approach to leverage phylogenetic analysis of metagenomic sequence data to conduct several types of analysis. First, we present a method to conduct phylogeny-driven Bayesian hypothesis tests for the presence of an organism in a sample. Second, we present a means to compare community structure across a collection of many samples and develop direct associations between the abundance of certain organisms and sample metadata. Third, we apply new tools to analyze the phylogenetic diversity of microbial communities and again demonstrate how this can be associated to sample metadata.
These analyses are implemented in an open source software pipeline called PhyloSift. As a pipeline, PhyloSift incorporates several other programs including LAST, HMMER, and pplacer to automate phylogenetic analysis of protein coding and RNA sequences in metagenomic datasets generated by modern sequencing platforms (e.g., Illumina, 454).
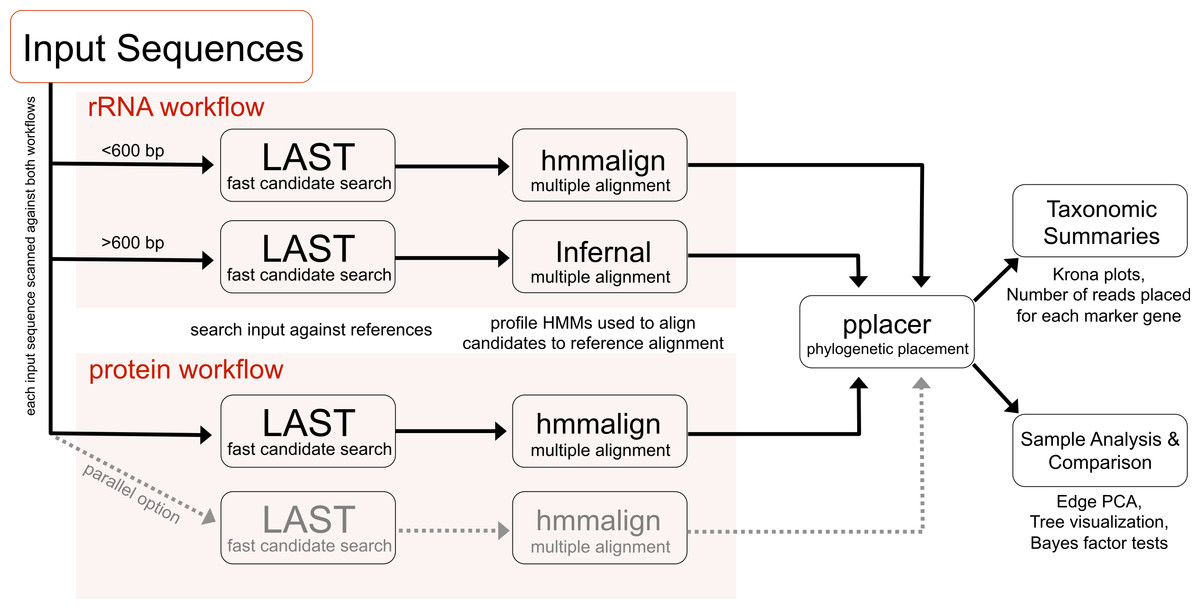 |
| Figure 1 showing the Phylosift workflow. |
The workflow follows a series of steps including
- Sequence identity search
- Alignment to reference multiple alignment
- Placement on a phylogenetic reference tree
- Visual presentation of taxonomic summary
- Comparison among samples (e.g., using Edge PCA)
- Acquiring new genome data
- Gene family search and alignment workflow on each genome
- Phylogenetic inference and pruning
- Selection of representatives for similarity search
- Taxonomic reconciliation
The paper shows some of the things you can do with Phylosift and some comparison of Phylosift and other methods.
 |
| Figure 2. Comparison of QIIME PCA and edge PCA analysis of human fecal samples. |
 |
| Figure 3: Lineages contributing variation in human fecal sample community structure. (Analyzed using EDGE PCA) |
It also provides Krona based output visualization of the taxonomic composition of a sample.
Anyway, more on Phylosift later. Just thought I would get some out here on the blog. Thanks to Aaron Darling, Holly Bik, Guillaume Jospin, Eric Lowe and Erick Matsen for all their hard work on this. And thanks to the Department of Homeland Security for supporting the work.
For more about Phylosift see
Worth a look: American Academy of Microbiology report on the Human Microbiome
Definitely worth checking this out: FAQ: Human Microbiome, January 2014. It is a report from the American Academy of Microbiology and it is really well done. In addition to the report itself there is also and Infographic and a nice little handout.
The report was based on discussions with a collection of Human Microbiome Gurus:
And it was written by Ann Reid and Shannon Greene. It has a variety of useful tidbits and has a reasonable number of caveats – such as “it should be noted, however, that at this point, most studies, even in mice, are looking at correlations between gut microbiome composition and factors like weight, insulin sensitivity, and other metabolic measures.”
#UCDavis encouraging those with the flu to stay home rest and recover
Good to see this. I note – I have a lab policy that anyone who is sick MUST stay home. That forced my home for most of this week despite actually wanting to be in the lab / office for work.
See forwarded email below:
Dear MSOs/CAOs,
Following up on recent news reports about the major flu virus in the Sacramento area, Human Resources has reminded us to please encourage those are sick to please stay home, rest and recover. This will help prevent colleagues and friends from getting the flu.
While it takes about two weeks for the flu shot to work, it is probably not too late to get one since there are least two more months of flu season.
Please circulate this information us as is appropriate in your department/center.
Thanks.
Donna
Donna Watkins Olsson
Executive Assistant Dean

New EisenLab paper: PhyloSift: phylogenetic analysis of genomes and metagenomes [PeerJ]
New paper from people in the Eisen lab (and some others): PhyloSift: phylogenetic analysis of genomes and metagenomes [PeerJ]. This project was coordinated by Aaron Darling, who was a Project Scientist in my lab and is now a Professor at the University of Technology Sydney. Also involved were Holly Bik (post doc in the lab), Guillaume Jospin (Bioinformatics Engineer in the lab), Eric Lowe (was a UC Davis undergrad working in the lab) and Erick Matsen (from the FHCRC). This work was supported by a grant from the Department of Homeland Security.
Abstract:
Like all organisms on the planet, environmental microbes are subject to the forces of molecular evolution. Metagenomic sequencing provides a means to access the DNA sequence of uncultured microbes. By combining DNA sequencing of microbial communities with evolutionary modeling and phylogenetic analysis we might obtain new insights into microbiology and also provide a basis for practical tools such as forensic pathogen detection.
In this work we present an approach to leverage phylogenetic analysis of metagenomic sequence data to conduct several types of analysis. First, we present a method to conduct phylogeny-driven Bayesian hypothesis tests for the presence of an organism in a sample. Second, we present a means to compare community structure across a collection of many samples and develop direct associations between the abundance of certain organisms and sample metadata. Third, we apply new tools to analyze the phylogenetic diversity of microbial communities and again demonstrate how this can be associated to sample metadata.
These analyses are implemented in an open source software pipeline called PhyloSift. As a pipeline, PhyloSift incorporates several other programs including LAST, HMMER, and pplacer to automate phylogenetic analysis of protein coding and RNA sequences in metagenomic datasets generated by modern sequencing platforms (e.g., Illumina, 454).
For more about Phylosift see
#UCDavis EVE Ecologist search candidate Stephanie Yelenik – 1/9
DEPARTMENT OF EVOLUTION AND ECOLOGY
RECRUITMENT SEMINAR
ECOLOGIST
Dr. Stephanie Yelenik
Research Ecologist
U.S. Geological Survey
Pacific Islands Ecosystem Research Center
“Plant-soil interactions, community dynamics and ecosystem management"
Thursday, January 9th, 2014
2:10pm
1022 Life Sciences Building
Faculty Host: Professor Gail Patricelli, Department of Evolution and Ecology







