Category: Misc.
Wrap up of tweets from Joe Derisi talk
Just a quick post here – for those not following on Twitter – Joe Derisi gave a talk at UC Davis 1/9 and I posted some tweets about it. Here they are:
| phylogenomics Joe DeRisi getting ready for his talk at the #ucdavis Genome Center this am http://t.co/OwZbM2nn 1/9/12 9:57 AM |
| phylogenomics Joe Derisi’s first slide : “Bees, viruses, and plastids: a seminar in two parts” – I think he needs to work on his math http://t.co/0nPjChlc 1/9/12 10:05 AM |
| phylogenomics DeRisi works with mobile honeybee colony trucks that travel around the country to service pollination needs 1/9/12 10:09 AM |
| phylogenomics DeRisi – one of the problems with figuring out what is causing CCD is we do not know much about “normal” healthy colonies 1/9/12 10:09 AM |
| phylogenomics Bee trucks start in Mississippi in winter, spring in Dakotas, then To california for almonds 1/9/12 10:10 AM |
| phylogenomics DeRisi got into bee studies because of colony collapse disorder CCD 1/9/12 10:13 AM |
| phylogenomics DeRisi – grinds up bees and then assays them with “bee pathogen chip” and sequencing 1/9/12 10:13 AM |
| phylogenomics Derisi discussing his new Plos one w/ SFSU paper showing phorid flies associated with bees 1/9/12 10:16 AM |
| phylogenomics Derisi says if you want your paper to get 1000s of hits you should mention Zombies somewhere as they did w/ “Zombie bees” 1/9/12 10:18 AM |
| phylogenomics Derisi: zombie bees are cool but the phorids are probably not associated with CCD 1/9/12 10:19 AM |
| phylogenomics DeRisi found six major pathogens associated w/ bees: Nosema, phorids, spiroplasma, notovirus, and others 1/9/12 10:22 AM |
| phylogenomics DeRisi – sequencing ground up bees – most reads are host or pathogens they know about – says key to characterizing new viruses is assembly 1/9/12 10:23 AM |
| phylogenomics DeRisi mentioning their PRICE assembler – an inductive assembly method 1/9/12 10:25 AM |
| notSoJunkDNA @phylogenomics URL for PRICE is http://t.co/bRGiY7Rz 1/9/12 10:30 AM |
| phylogenomics Why I love Joe Derisi “there will be a manuscript on the assembler someday… But the software is available for free now” 1/9/12 10:27 AM |
| phylogenomics DeRisi pooled together Bee samples prepped with various methods so that he could get diverse viruses covered 1/9/12 10:29 AM |
| phylogenomics DeRisi – lake sinai virus 2 is present in up to 10^11 copies per bee 1/9/12 10:35 AM |
| phylogenomics DeRisi – also present in bees – Crithidia – a trypanosome – related to known pathogens of other insects 1/9/12 10:36 AM |
| phylogenomics DeRisi – Crithidia also increases in abundance in winter 1/9/12 10:36 AM |
| phylogenomics DeRisi – sequenced and assembled genome of Crithidia – genome is interesting 1/9/12 10:37 AM |
| phylogenomics DeRisi – part two – the essential function of the Plasmodium apicoplast 1/9/12 10:38 AM |
| phylogenomics DeRisi – history of Studies of plasmodium apicoplast – organelle bound by four membranes – likely b/c result of secondary symbiosis 1/9/12 10:39 AM |
| phylogenomics Derisi discussing sequencing of apicoplast genome in 1990s by Gardner, McFadden, etc 1/9/12 10:40 AM |
| phylogenomics DeRisi – experiments suggest IPP is the sole essential product of apicoplast biology 1/9/12 10:47 AM |
Draft post cleanup #13: Twisted tree of life award: MSNBC, Aliens and Photosynthesis
Yet another post in my “draft blog post cleanup” series. Here is #13 from July 2010.
I wanted to give this article a “Twisted Tree of Life Award“:
How to find aliens: Follow the photosynthesis – Technology & science – Space – Space.com – msnbc.com
It is pretty painful to me. Basically the people they are quoting argue that since “complex” life on Earth requires oxygen and since oxygen only comes from photosynthesis, therefore we should look for planets where photosynthesis is possible as the place where life is most likely to be interesting. Uggh. So many things in the article I did not like … but just not enough time I guess to bitch about it then. I will leave it to readers to decide for themselves I guess …
To be discussed in journal club here today: eukaryotic metatranscriptomics
Going to be discussing this in a journal club today
Note – am copying the whole text here to mark it up a bit since it it awkward to try and mark it up at the PLoS One site
Metatranscriptomics Reveals the Diversity of Genes Expressed by Eukaryotes in Forest Soils
- To add a note, highlight some text. Hide notes
- Make a general comment
Abstract Top
Introduction Top
Results Top
Sequence datasets (Table 1)
Taxonomic diversity of the soil eukaryotic communities
doi:10.1371/journal.pone.0028967.g001
Global functional annotation of the cDNAs
doi:10.1371/journal.pone.0028967.t002
Targeted annotation of the cDNAs
doi:10.1371/journal.pone.0028967.t003
doi:10.1371/journal.pone.0028967.t004
Identification of full-length CAZymes
doi:10.1371/journal.pone.0028967.g002
Discussion Top
Materials and Methods Top
Study site and soil sampling
RNA extraction, cDNA libraries construction and sequencing
18S rDNA gene libraries construction and sequencing
cDNA sequences cleaning and clustering
Global cDNA sequences annotation
Targeted annotation of cDNAs
Taxonomic annotation of rRNA and cDNA sequences
Phylogenetic analyses
Supporting Information Top
Acknowledgments Top
Author Contributions Top
References Top
- (2008) Links between plant litter chemistry, species diversity, and below-ground ecosystem function. Proc Natl Acad Sci U S A 105: 19780–19785. FIND THIS ARTICLE ONLINE
- (2008) Plant species traits are the predominant control on litter decomposition rates within biomes worldwide. Ecol Lett 11: 1065–1071. FIND THIS ARTICLE ONLINE
- (2008) Stoichiometry of soil enzyme activity at global scale. Ecol Lett 11: 1252–1264. FIND THIS ARTICLE ONLINE
- (2004) Genome sequence of the lignocellulose degrading fungusPhanerochaete chrysosporium strain RP78. Nat Biotechnol 22: 695–700. FIND THIS ARTICLE ONLINE
- (2008) Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nat Biotechnol 26: 553–560. FIND THIS ARTICLE ONLINE
- (2009) Genome, transcriptome, and secretome analysis of wood decay fungus Postia placenta supports unique mechanisms of lignocellulose conversion. Proc Natl Acad Sci U S A 106: 1954–1959.FIND THIS ARTICLE ONLINE
- (2007) Genome sequence of the lignocellulose-bioconverting and xylose-fermenting yeast Pichia stipitis. Nat Biotechnol 25: 319–26.FIND THIS ARTICLE ONLINE
- (1998) Fungal succession – unravelling the unpredictable. Mycol Res 102: 1–15. FIND THIS ARTICLE ONLINE
- (2010) The influence of vegetation type, soil properties and precipitation on the composition of soil mite and microbial communities at the landscape scale. J Biogeogr 37: 1317–1328.FIND THIS ARTICLE ONLINE
- (2011) Contrasting diversity patterns of crenarchaeal, bacterial and fungal soil communities in an alpine landscape. PLoS ONE 6: e19950. FIND THIS ARTICLE ONLINE
- (2009) Gene-centric metagenomics of the fiber-adherent bovine rumen microbiome reveals forage specific glycoside hydrolases. Proc Natl Acad Sci U S A 106: 1948–1953. FIND THIS ARTICLE ONLINE
- (2011) Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331: 463–7. FIND THIS ARTICLE ONLINE
- (2011) Snapshot of the Eukaryotic gene expression in Muskoxen rumen – A metatranscriptomic approach. PLoS ONE 6: e20521. FIND THIS ARTICLE ONLINE
- (2010) Functional metagenomics to mine human gut microbiome for dietary fiber catabolic enzymes. Genome Res 20: 1605–1612. FIND THIS ARTICLE ONLINE
- (2009) Adaptation to herbivory by the Tammar wallaby includes bacterial and glycoside hydrolase profiles different from other herbivores. Proc Natl Acad Sci U S A 107: 14793–798. FIND THIS ARTICLE ONLINE
- (2009) Parallel metatranscriptome analyses of host and symbiont gene expression in the gut of the termite Reticulotermes flavipes. Biotechnol Biofuels 2: 25. FIND THIS ARTICLE ONLINE
- (2007) Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450: 560–5.FIND THIS ARTICLE ONLINE
- (2010) Targeted discovery of glycoside hydrolases from a switchgrass-adapted compost community. PLoS ONE 5: e8812.FIND THIS ARTICLE ONLINE
- (2006) Identification of eukaryotic open reading frames in metagenomic cDNA libraries made from environmental samples. Appl Environ Microbiol 72: 135–143. FIND THIS ARTICLE ONLINE
- (2007) Soil eukaryotic functional diversity, a metatranscriptomic approach. ISME J 1: 632–642. FIND THIS ARTICLE ONLINE
- (2008) Tracking microbial biodiversity through molecular and genomic ecology. Res Microbiol 159: 67–73.FIND THIS ARTICLE ONLINE
- (2008) Environmental rRNA inventories miss over half of protistan diversity. BMC Microbiol 8: 222. FIND THIS ARTICLE ONLINE
- (2011) Myxomycetes in soil. Soil Biol Biochem 43: 2237–2242. FIND THIS ARTICLE ONLINE
- (2010) MetaBioME: a database to explore commercially useful enzymes in metagenomics datasets. Nucl Ac Res 38: D468–D472. FIND THIS ARTICLE ONLINE
- (2007) The cytochrome P450 engineering database: a navigation and prediction tool for the cytochrome P450 protein family. Bioinformatics 23: 2015–2017. FIND THIS ARTICLE ONLINE
- (2006) TCDB: the transporter Classification Database for membrane transport protein analyses and information. Nucl Acids Res 34: D181–D186. FIND THIS ARTICLE ONLINE
- (2010) Multiple lateral gene transfers and duplications have promoted plant parasitism ability in nematodes. Proc Natl Acad Sci U S A 107: 17651–17656. FIND THIS ARTICLE ONLINE
- (2006) Reconstructing the early evolution of Fungi using a six-gene phylogeny. Nature 443: 818–22. FIND THIS ARTICLE ONLINE
- (2011) Insight into oxidative degradation of cellulose by a copper metalloenzyme that exploits biomass components. Proc Natl Acad Sci U S A 108: 15079–15084. FIND THIS ARTICLE ONLINE
- (2010) Characterization of the endoglucanase and glucomannanase activities of a glycoside hydrolase family 45 protein from Penicillium decumbens 114-2. J Gen Appl Microbiol 56: 223–229. FIND THIS ARTICLE ONLINE
- (2009) Molecular cloning of glycoside hydrolase family 45 cellulase genes from brackish water clamCorbicula japonica. Comp Biochem Physiol B Biochem Mol Biol 152: 390–396. FIND THIS ARTICLE ONLINE
- (1994) A novel, small endoglucanase gene, egl5, from Trichoderma reesei isolated by expression in yeast. Mol Microbiol 13: 219–228. FIND THIS ARTICLE ONLINE
- (2000) Purification, characterization and amino-acid sequence analysis of a thermostable, low molecular mass endo-ß-1,4-glucanase from blue mussel, Mytilus edulis. Eur J Biochem 267: 4970–4977. FIND THIS ARTICLE ONLINE
- (2008) Microbial community gene expression in ocean surface waters. Proc Natl Acad Sci U S A 105: 3805–3810. FIND THIS ARTICLE ONLINE
- (2008) Detection of large numbers of novel sequences in the metatranscriptomes of complex marine microbial communities. PLoS ONE 3: e3042. FIND THIS ARTICLE ONLINE
- (2010) The taxonomic and functional diversity of microbes at a temperate coastal site: a ‘multi-omic’ study of seasonal and diel temporal variation. PLoS ONE 5: e15545. FIND THIS ARTICLE ONLINE
- (2009) Transcriptional activity of paddy soil bacterial communities. Environ Microbiol 11: 960–970. FIND THIS ARTICLE ONLINE
- (2008) Simultaneous assessment of soil microbial community structure and function through analysis of the metatranscriptome. PLoS ONE 3: e2527. FIND THIS ARTICLE ONLINE
- (2010) Validation of two ribosomal RNA removal methods for microbial metatranscriptomics. Nature Met 10: 807–812. FIND THIS ARTICLE ONLINE
- (2010) Development and quantitative analyses of a universal rRNA-substraction protocol for microbial metatranscriptomics. ISME J 4: 896–907. FIND THIS ARTICLE ONLINE
- (1997) Polyadenylation of mRNA in prokaryotes. Ann Rev Biochem 66: 173–197. FIND THIS ARTICLE ONLINE
- (2011) Genome sequencing and comparative transcriptomics of the model entomopathogenic fungi Metarhizium anisopliae and M. acridum. PLoS Genet 7: e1001264. FIND THIS ARTICLE ONLINE
- (2010) Périgord black truffle genome uncovers evolutionary origins and mechanisms of symbiosis. Nature 464: 1033–1038. FIND THIS ARTICLE ONLINE
- (2010) Genome sequences of the human body louse and its primary endosymbiont provide insights into the permanent parasitic lifestyle. Proc Natl Acad Sci U S A 107: 12168–12173. FIND THIS ARTICLE ONLINE
- (2011) The ecoresponsive genome of Daphnia pulex. Science 331: 555–561. FIND THIS ARTICLE ONLINE
- (2011) A novel fungal family of oligopeptide transporters identified by functional metatranscriptomics of soil eukaryotes. ISME J. doi:10.1038/ismej.2011.67.
- (2009) Plant cell walls throughout evolution: towards a molecular understanding of their design principles. J Exp Bot 60: 3615–3635. FIND THIS ARTICLE ONLINE
- (2010) Quantification of monosaccharide composition of hemicelluloses from different plant functional types. Plant Physiol Biochem 48: 1–8. FIND THIS ARTICLE ONLINE
- (2006) Effect of tree species substitution on organic matter biodegradability and mineral nutrient availability in a temperate topsoil. Ann For Sci 63: 763–771. FIND THIS ARTICLE ONLINE
- (2010) Molecular insight into lignocellulose digestion by a marine isopod in the absence of gut microbes. Proc Natl Acad Sci U S A 107: 5345–5350. FIND THIS ARTICLE ONLINE
- (2010) Diversity of beetle genes encoding novel plant cell wall degrading enzymes. PLoS ONE 5: e15635. FIND THIS ARTICLE ONLINE
- (2010) Screening of a soil metatranscriptomic library by functional complementation ofSaccharomyces cerevisiae mutants. Microbiol Res 166: 360–368.FIND THIS ARTICLE ONLINE
- (2011) Activity-based metagenomic screening and biochemical characterization of bovine ruminal protozoan glycoside hydrolases. Appl Environ Microbiol. doi:10.1128/AEM.05925-11.
- (2010) Performance of the COX1 gene as a marker for the study of metabolically active Pezizomycotina and Agaricomycetes fungal communities from the analysis of soil RNA. FEMS Microbiol Ecol 74: 693–705. FIND THIS ARTICLE ONLINE
- (2001) Study of genetic diversity of eukaryotic picoplankton in different oceanic regions by small-subunit rRNA gene cloning and sequencing. Appl Environ Microbiol 67: 2932–2941. FIND THIS ARTICLE ONLINE
- (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22: 1658–1659. FIND THIS ARTICLE ONLINE
- (2005) Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21: 3674–367. FIND THIS ARTICLE ONLINE
- (2000) Gene ontology: tool for the unification of biology. Nat Genet 25: 25–29. FIND THIS ARTICLE ONLINE
- (2009) The Carbohydrate-Active EnZyme database (CAZy): an expert resource for glycogenomics. Nucl Ac Res 37: D233–D238.FIND THIS ARTICLE ONLINE
- (2008) FOLy: an integrated database for the classification and functional annotation of fungal oxidoreductases potentially involved in the degradation of lignin and related aromatic compounds. Fung Genet Biol 45: 638–645. FIND THIS ARTICLE ONLINE
- (2009) The cytochrome P450 Engineering Database: integration of biochemical properties. BMC Biochem 10: 27. FIND THIS ARTICLE ONLINE
- (2003) The lipase engineering database: a navigation and analysis tool for protein families. Nucl Ac Res 31: 319–321. FIND THIS ARTICLE ONLINE
- (2009) MEROPS: the peptidase database. Nucl Ac Res 38: D227–D233. FIND THIS ARTICLE ONLINE
- (2010) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27: 221–224. FIND THIS ARTICLE ONLINE
- (2009) Methods for comparative metagenomics. BMC Bioinformatics 10: Suppl 1S12. FIND THIS ARTICLE ONLINE
- (2008) Phylogeny.fr: a robust phylogenetic analysis for the non-specialist. Nucl Acids Res 36: W465–W469. FIND THIS ARTICLE ONLINE
- (2009) Post-genomic insight into the plant polysaccharide degradation potential of Aspergillus nidulans and comparison toAspergillus niger and Aspergillus oryzae. Fung Genet Biol 46: S161–S169. FIND THIS ARTICLE ONLINE
Draft post cleanup #12: RecA is cool (and interesting)
Yet another post in my “draft blog post cleanup” series. Here is #12:
I have been interested in RecA and related proteins for many many years. In particular I have been interested in structural and functional evolution of RecA and its relatives. This all started when for my second scientific paper I helped a post doc in the lab where I was doing my PhD do some structure-function-evolution studies (with a little help from Chris Lee, then in Mike Levitt’s lab, and my brother, then in Don Wiley’s lab).
For my first talk at a scientific meeting I discussed using RecA as a marker for phylogenetic studies (and had a slide where I had text saying RecA was cool). Over the years I have continued to try and study RecA or at least use it for studying microbial diversity in some way. See for example
- The RecA protein as a model molecule for molecular systematic studies of bacteria: comparison of trees of RecAs and 16S rRNAs from the same species
- The phylogenetic relationships of Chlorobium tepidum and Chloroflexus aurantiacus based upon their RecA sequences
- Analysis of Deinococcus radiodurans’s transcriptional response to ionizing radiation and desiccation reveals novel proteins that contribute to extreme radio resistance
- Sequence characterization and comparative analysis of three plasmids isolated from environmental Vibrio spp.
- Stalking the fourth domain in metagenomic data: searching for, discovering, and interpreting novel, deep branches in marker gene phylogenetic trees)
Anyway – in this context I was excited to see a new paper on RecA-structure-function-evolution studies: PLoS Genetics: Separation of Recombination and SOS Response in Escherichia coli RecA Suggests LexA Interaction Sites from Olivier Lichtarge and others. In the paper they use Lichtarge’s Evolutionary Trace method to study RecA.
The paper is worth a look and if you are interested in structure-function-evolution types of studies and need a good protein to work on, I would suggest you look at RecA and its relatives … They are Cool.
Oh and for the fun of it — I have found some of my slides from that talk in 1995. Here they are
AAAS meeting – is this one for embargo watch?
Giving a talk at the AAAS meeting in February in Vancouver. I have avoided AAAS meetings previously because I do not like AAAS’s position on open access issues. Given that AAAS is at least indirectly a supporter of the recent Research Works Act I am pondering whether or not I will boycott the meeting. While I ponder that — I thought I would share the presenter instructions I just got from AAAS (see below).
Apparently, my talk is “embargoed” – though I am not sure I understand how that works for a talk (see the part I highlighted in yellow which, well, I almost certainly will not be following). I do not understand actually what a talk embargo means – am I supposed to not share with people what I am working on so that every piece of data I present at the meeting will never have ben seen by anyone? Or am I just not supposed to show my talk to anyone? What exactly is a talk embargo? And what will they do when I do not follow it? Maybe Ivan Oransky knows.
I note – I am surprised AAAS does not try to require me to sign over rights to my presentation to them …
This request for materials is from the AAAS media relations team and is separate from any you may receive from your symposium organizer or the AAAS Annual Meeting office.
—————————————-
Dear AAAS Annual Meeting Participant:
If you have already uploaded your materials to the Virtual Newsroom for the 2012 AAAS Annual Meeting in Vancouver, thank you and please disregard the rest of this e-mail.
For those speakers who have not submitted materials, we’d appreciate your prompt attention to this request. We expect a good turnout of reporters at the meeting in February, and we’d like to provide them as much information as possible about your presentation.
Symposium organizers can help as well by uploading relevant papers or overview documents and encouraging your speakers to submit materials. Papers and speaker materials are for use by reporters in preparing stories and are not made available to general registrants at the meeting.
Speakers and organizers can submit materials by going to:
http://www.eurekalert.org/aaasnewsroom/mcm/speakersYour individual username and password for the site:
Username: xxxx
Password: xxxxPlease provide the following:
— A one-paragraph biographical sketch (not a C.V.)
— A short lay-language summary of your talk, beyond the abstract.
— The text of your talk, if available, or a related (ideally recent) technical paper, either as a Word file or a PDF. PowerPoint presentations are acceptable, but a full text will better serve reporters’ needs.
— Any additional supporting materials, including multimedia files such as JPEG or TIFF photos in high resolution (300 dpi) and/or digitized video clips.
IMPORTANT: Please note that all AAAS meeting presentations are strictly embargoed and your speaker materials should not be released publicly until the time of your presentation.
If you upload your materials by 16 January, we will copy them at our expense for placement in the on-site library of speaker materials, available only to newsroom registrants.
Please notify your institution’s press office of your AAAS Annual Meeting presentation as soon as possible. Your press office can help you submit speaker materials to us and can begin to generate media interest.
If you have any questions, please feel free to contact us.
Cracking the microbial code: Pam Ronald @pcronald and her story behind recent papers
Pam Ronald asked me to retract this post because she discovered that one of the strains used in the reported experiments was compromised. She notes that he laboratory is now in the process of repeating each experiment with newly validated strains and that much of the work has been independently validated in other laboratories (McCarthy et al., 2011, J Bacteriology 193:6375-6378; Shuguo et al. 2012, Appl Biochem Biotechnol. 166:1368-79).
Please contact her if you have questions. I am leaving the text of the post below based on the notion that one should not completely delete anything from the record but rather post corrections and retractions. More detail will be coming from Pam soon. Kudos to Pam for trying to make sure the scientific record is accurate and for contacting me about this.

Very very excited for this “Story behind the paper” post. For those who do not know, I have been hosting posts here on my blog written by authors of Open Access papers telling the story behind the paper (I have also been writing some of my own). This one comes from my friend and UC Davis colleague Pamela Ronald. Pam is a fascinating person – great scientist – fun in person – blogger – book author – prize winner for work for the developing world – and much more. She was in the office across the hall from me for a while but now has moved to new diggs on campus and I miss the regular interactions with her.Anyway – without further ado – here is her post describing a new paper of hers from PLoS One, among many other things (note – she wrote the post – I added a few headers and pictures and links). Finally I note – if anyone else wants to tell the Story behind one of their papers – please let me know – I would love to host it here.
 |
The windowless room, dank an dark, was not an obvious place for inspiration. I took notes, wondering if I would be able to glean anything meaningful from Professor Helen Stafford’s (1922-2011) meandering lecture. I was skeptical. After all, this was the same teacher who, annoyed with our choice of vegetarianism, had told us that “plants have feelings, too”.
 |
In 1905, British geneticist and plant breeder Rowland Biffen demonstrated that it as possible to generate wheat varieties with resistance to a devastating diseases by moving genes around. He cross-pollinated a resistant wheat variety with a susceptible wheat variety and showed that the resulting seed carried the resistance of the parent.
These were central questions addressed by the laboratory of Brian Staskwicz, where I carried out my PhD work. He had recently identified a bacterial protein that triggered an immune response in infected plants. He later identified one of the first plant resistance genes. After graduating, I moved to the laboratory of plant breeder Steven Tanksley at Cornell who was pioneering gene mapping in plants using molecular methods.Focusing on Xa21
 |
My focus was a genetic locus, called Xa21, described in 1977 by Drs. S. Devadath, Gurdev Khush and collaborators. Rice plants carrying Xa21 have an unusual property: they are resistant to all known races of the bacterial pathogen Xanthomonas oryzae pv. oryzae (Xoo), which normally caused a devastating disease of rice in Asia and Africa. In 1992, I mapped this locus to a specific region in the rice genome. I hypothesized that it must consist of a cluster of tightly linked genes each recognizing a single race of the pathogen or a single gene that encoded a receptor recognizing a conserved microbial signature present in all races. When I moved to UC Davis I began a map-based cloning approach to isolate Xa21.
A few years after the discovery of the first plant resistance genes, the fly Toll and mouse Toll-like receptor (Tlr4) genes were isolated and shown to have striking structural similarities to XA21. Like XA21, TLR4 is membrane bound extracellular receptor that was predicted to bind a conserved microbial signature. TLR4 also carry the Toll /IL-1 Receptor (TIR) domain found in fly TOLL, the tobacco N resistance gene and the flax L6 resistance gene.
Thus, the discovery of a role for Toll and TLR4 in immunity provided a structural link between sensors utilized by plants and animals to detect infection. Professors Bruce Beutler and Jules Hoffman were awarded the 2011 Nobel Prize in Physiology or Medicine for their important work.
Characterizing Ax21
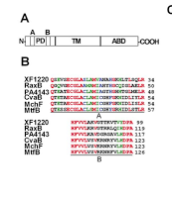 |
To isolate this molecule, which we named Ax21 (Activator of Xa21-mediated immunity), we screened for bacterial mutants altered in their ability to produce active Ax21. In this way, we isolated and characterized eight genes required for Ax21 activity (rax genes). raxA, raxB and raxC encode components of a predicted type I secretion system. Ax21 requires this RaxABC system for activity and secretion. The RaxB protein carries two highly conserved domains characteristic of proteins in Gram-positive bacteria that cleave N-terminal peptides prior to secretion of small proteins. Another set of rax genes encoded proteins important for sulfation.
 |
Over the last 20 years, researchers have shown that bacteria employ specific signals to communicate. These signaling molecules, called “bacterial Esperanto” by Professor Bonnie Bassler, an early pioneer in studies of bacterial communication, accumulate in the external environment as the cells grow. When the concentration reaches a certain threshold level, the bacteria mobilize together to carry out concerted, group actions. This process is called quorum sensing.
Most rice plants are virtually defenseless against this Ax21-mediated bacterial attack – except for those plants that carry the XA21 immune receptor. This early detection gives the plant time to mobilize its defenses and mount an early and potent immune response.
 |
The discovery that a small protein from a Gram-negative bacterium has a dual role in bacterial communication and in activation of the host innate immune response has not previously been demonstrated. We do not, however, believe this is an anomaly or that the biological importance of Ax21 is restricted to plant pathogens. We previously reported that Ax21 is also conserved in the nosocomial pathogen S. maltophilia and proposed a similar role for Ax21 in this species. Consistent with our hypothesis, a synthetic Ax21 protein has now been shown to regulate gene expression, motility, and biofilm formation in S. maltophilia, extending our findings to an animal pathogen.

![]()
Han, S., Sriariyanun, M., Lee, S., Sharma, M., Bahar, O., Bower, Z., & Ronald, P. (2011). Small Protein-Mediated Quorum Sensing in a Gram-Negative Bacterium PLoS ONE, 6 (12) DOI: 10.1371/journal.pone.0029192
Press release from De Gruyter regarding acquisition of Versita re #OpenAccess
This press release may be of interest to those interested in Open Access publishing so I am posting it here:
De Gruyter acquires Versita and becomes third-biggest international Open Access publisher
As of 2012 De Gruyter will be merging the newly acquired Open Access journals with its traditional subscription-based and freely sold content on one electronic platform, thus providing researchers with an outstanding service: users will have a powerful search interface for searching all academic articles from journals, books and databanks in which the articles have been published, independently of the various business models.
Draft post cleanup #11: Tree Hugging
Yet another post in my “draft blog post cleanup” series. Here is #11 from September.
Just a quick one. In August a nice review paper came out on phylogenetic analysis software: Learning to Become a Tree Hugger | The Scientist. By Amy Maxmen it is a “A guide to free software for constructing and assessing species relationships”. Definitely worth checking out.
Draft post cleanup #10: trip to LA artificially sweetened by Carolyn de la Peña
Yet another post in my “draft blog post cleanup” series. Here is #10:
Went on a mini trip to UCLA for a mini meeting in November. It seemed appropriate that I brought with me to Los Angeles, land of empty pleasures – the new book from UC Davis Professor Carolyn de la Peña – “Empty Pleasures” on the history of artificial sweeteners. So I took a picture of the book overlooking part of LA from my hotel room:
The book is great read by the way …



















