A couple of people have requested that I post information on the various kits and probes being used to assay water chemistry for our aquarium study. Here’s the list, divided by type with a link to each item on Amazon. Also various user complaints.
Probes:
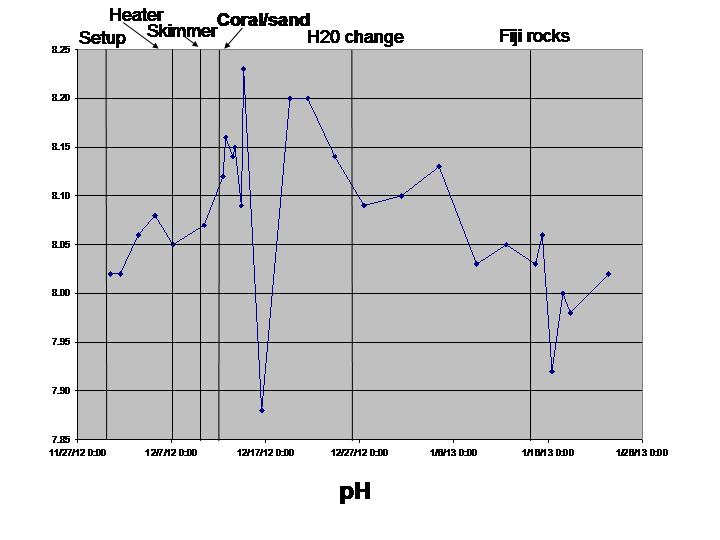
pH (worked fine, held calibration well)
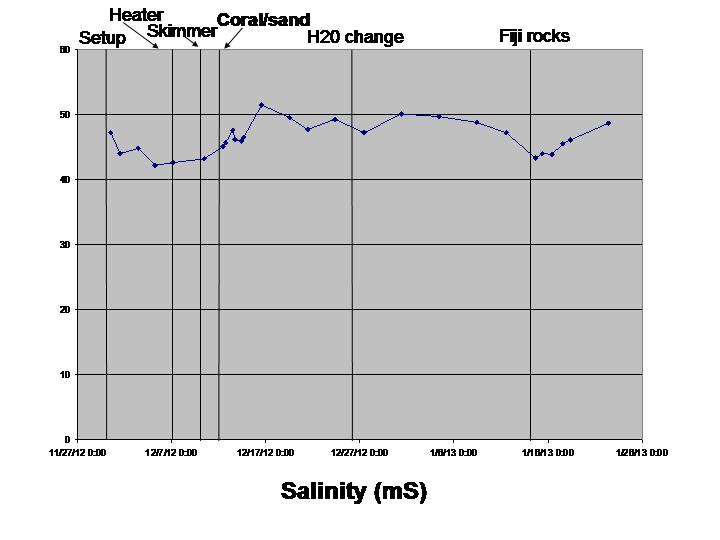
Salinity (worked fine, held calibration well)
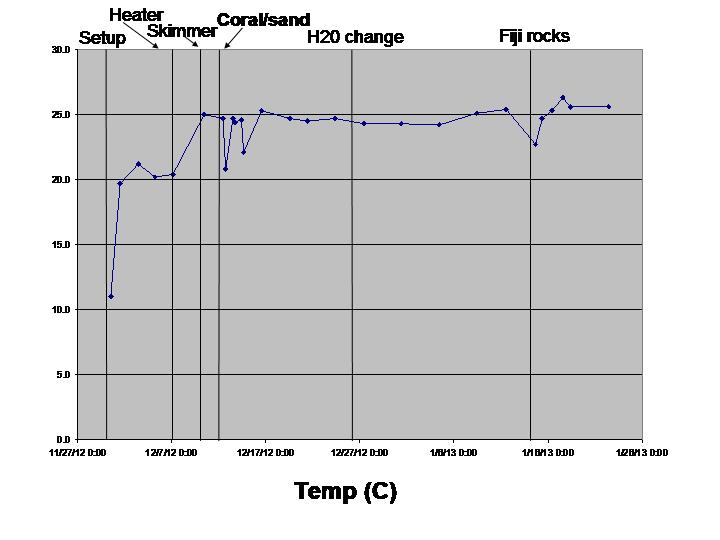
Temperature (annoying, a bit fiddly, I wouldn’t do this one again)
Titration-based kits:
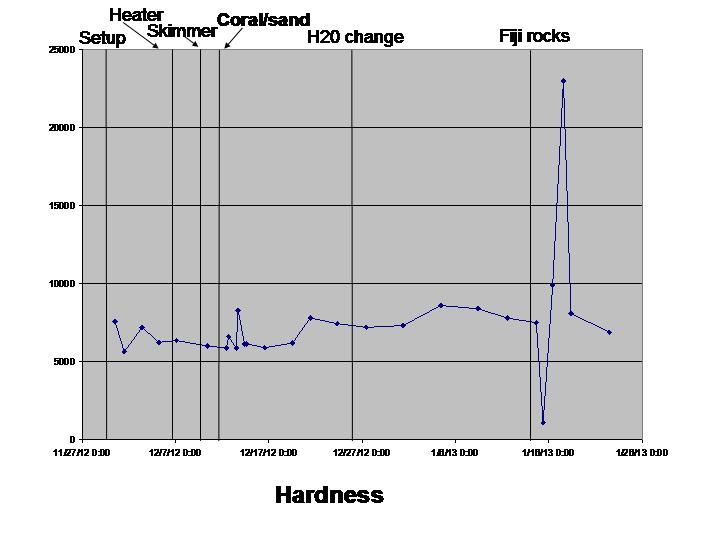
Hardness, Alkalinity, Chloride, and Sulfide were all measured using this combo package It also includes an iron assay that we didn’t use. All of these kits are prone to error since they’re titration based. But used very carefully (takes time!) they seemed to produce okay results. I recommend using glass flasks instead of the plastic beakers supplied.
Colorimetric scanners:
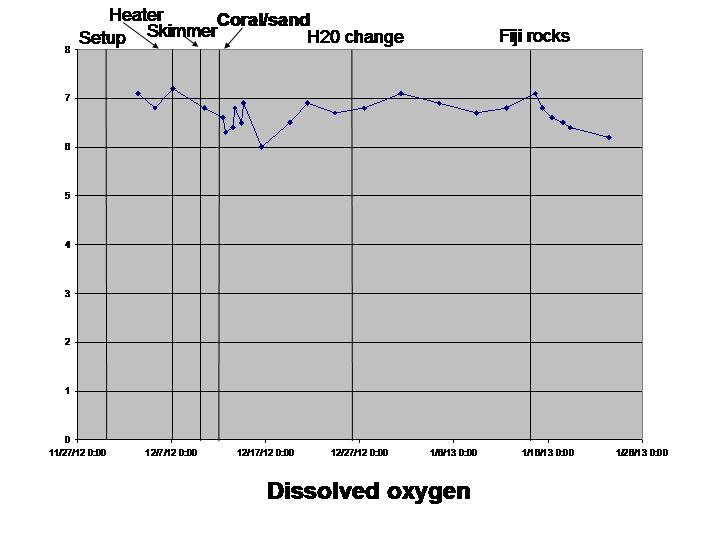
Dissolved oxygen (this kit requires that you have a glass container capable of holding exactly 60 mls of water.. they don’t tell you this until you read the instructions. Otherwise worked fine. A bit hard to use but conversely you’ll learn new vocabulary from reading the instructions)
Ammonia (worked fine, but you have to be very careful to follow the instructions and mix between adding reagents or you’ll get a false high reading)
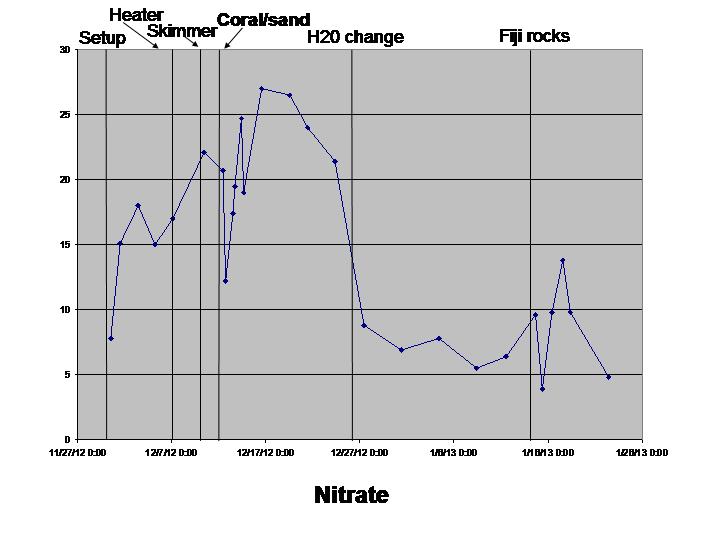
Nitrate (worked fine)
Nitrite (this one is pretty annoying; it’s hard to get the reagents into the tiny vial and these handheld meters turn themselves off after only 2 minute so if you get distracted you have to start over)
Phosphorus (see nitrite, but even worse. As far as we can tell it’s not even possible to follow the instructions for this since it turns itself off before you finish mixing the reagents)