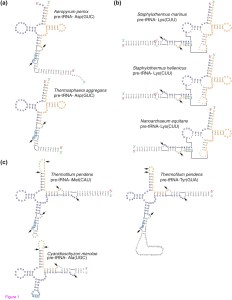
I was woefully unaware of some of the tRNA shenanigans going on in microbes until reading this paper: Genome Biology | Abstract | Discovery of permuted and recently split transfer RNAs in Archaea from Patricia Chan, Aaron Cozen and Todd Lowe. Life is pretty weird and wacky sometimes, even in components of cells that are considered “core” parts of the machinery of life. Go figure. It is worth a read …
Eisen Lab Blog
Hospital acquired infections – NY Times raising level of attention w/ editorial
Just a quick one here. The New York Times had an editorial Sunday on hospital acquired infections: Hospitals Shouldn’t Make You Sicker. I couldn’t agree more. The Times discussed some recent studies published in the New England Journal of Medicine as well as plans by the Obama Administration to fund some work in this area. It would be good I note if some of the funding focused not on infections per se but on the microbial ecology of hospitals, which we know very little about. Focusing just on hand washing and transmission of nasty pathogens is not enough.
Boston, Bioinformatics & Ben Franklin Award wrap up from #BioIT11
 |
| Photo by Mark Gabrenya |
Well, just got back from Boston where I went to the BioIT World convention to pick up the “Benjamin Franklin Award” for contributions to Open Science from Bioinformatics.Org. A quick round up of the trip:
Dropped off my stuff at the Seaport Hotel – had a nice view from my room.
 |
| Photo by Ashlee Earl |
 |
| Photo by Ashlee Earl |
Took the T back to the hotel after the game. And went to sleep. Got up very early to think about my “acceptance speech” for when I was to pick up my Ben Franklin Award. I made some quick slides on my Ipad (this was the first time I have gone to a meeting w/o my laptop) and during the talk before the award ceremony I emailed them to one of the organizers and we got things set up.
Then I was Introduced and Jeff Bizzaro read a mini statement about why I won the award. Something like what they put on the Bioinformatics.org web site:
Jonathan uses his high visibility in social media to advocate for open access by sharing links to discussions, mentioning open access articles and initiatives, and pushing for the opening up of popular closed access articles. This culture is shared with his students, who advocate for “open access” peer reviewing and created a peer-to-peer service for sharing bioinformatics material (articles, software and datasets). He is the academic editor in chief of PLoS Biology [1] and voices his opinions and support for open access publication and open data sharing on his “Tree of Life” blog [2]. In addition to just voicing his opinion, he also practices what he preaches, by refusing to publish in non-open access journals. With respect to bioinformatics, he has been involved with many software packages that are freely available, such as the recent AMPHORA [3] and PhyloOTU [4]. Lastly, Jonathan helped release a new open data sharing tool for scientists called BioTorrents [5]. This is just another step in encouraging all scientists to share their data and results more openly.
References:
4. http://www.ploscompbiol.org/article/info%3Adoi%2F10.1371%2Fjournal.pcbi.1001061
5. http://www.plosone.org/article/info:doi/10.1371/journal.pone.0010071
Note – am proud to get this award. It is given for contributions to Open Science and previous winners are an esteemed crew: Michael Eisen (my brother), Alex Bateman, Michael Ashburner, Jim Kent, Robert Gentleman, Phil Bourne, Lincoln Stein, Ewan Birney, and Sean Eddy (see full details here).
Then I gave my mini talk focusing on a brief history of how I got into Open Science. Here are my slides
Note the awkward typo where I introduced the “Public Library of Science”. Oops. Anyway – talked for a few minutes. While wearing my RedSox PLoS 1 shirt I note.
 |
| Photo by Jeff Bizarro |
 |
| Photo by Mark Gabrenya |
 |
| Photo by Mark Gabrenya |
And then there was a break. They took some pictures during the break and eventually I wandered around to the booths.
 |
| Photo by Mark Gabrenya |
 |
| Photo by Mark Gabrenya |
 |
| Photo by Mark Gabrenya |
I saw Nat Pearson who now works for Knome and I went to lunch with him to discuss my “Exome” which Knome has sequenced. I note, Nat was a student in a class I TAd at Stanford — good to see how far he has come.
And then back to the meeting where I wandered around again for a while. Saw an old friend from TIGR Xiaoying Lin who now works at Life Technologies and discussed the Ion Torrent with him.
Was pleased to see a booth giving away free RedSox tickets as a prize.
Then I headed out to Brookline for dinner with my Aunt and Uncle and cousin and eventually made my way back to the hotel where I had a few drinks.
The next day I got up a bit late, and eventually made my way to Logan Airport where the trip home was a disaster. My outbound flight was late. Missed my connection. Then the flight I was on was held up for others to make their connection. Though I did get a few hours in Denver Airport to wander around. Got home after 1 AM … And finally made it home.
Interesting PLoS One paper on local assembly from short reads by "tagging" DNA via restriction enzymes
Here is the abstract:
“Despite the power of massively parallel sequencing platforms, a drawback is the short length of the sequence reads produced. We demonstrate that short reads can be locally assembled into longer contigs using paired-end sequencing of restriction-site associatedDNA (RAD-PE) fragments. We use this RAD-PE contig approach to identify single nucleotide polymorphisms (SNPs) and determine haplotype structure in threespine stickleback and to sequence E. coli and stickleback genomic DNA with overlapping contigs of several hundred nucleotides. We also demonstrate that adding a circularization step allows the local assembly of contigs up to 5 kilobases (kb) in length. The ease of assembly and accuracy of the individual contigs produced from each RAD site sequence suggests RAD-PE sequencing is a useful way to convert genome-wide short reads into individually-assembled sequences hundreds or thousands of nucleotides long.”
Note as they note in the paper “Competing interests: E.A.J. has patents filed on the RAD marker, and partial interest in a company commercializing the system. This does not alter the authors’ adherence to all the PLoS ONE policies on sharing data and material” This seems like it would have potential in metagenomic applications. I note, we are working on a similar approach – and kind of got scooped here in a way. Hope their patent does not limit what we can do.
Hmm – How did I miss this paper discussing RFAM, Wikipedia & community annotation #cool
Very interesting discussion in this paper on Rfam about community annotation: Rfam: Wikipedia, clans and the “decimal” release. In the paper the authors (inlcuding Alex Bateman, Sean Eddy, Paul Gardner and others) discuss the use of Wikipedia for Community Annotation of biological databases. They report:
Given our positive experiences, we can highly recommend other curation efforts turning to Wikipedia for their annotation
We look forward to working with the wider community to develop these new tools and techniques.
It seems that they have bought into the Wikipedia based annotation system as having enormous potential. I generally agree though I am not sure how this is best done.
Nice review/commentary on challenges in phylogenomic analysis in PLoS Bio by Philippe et al. #fb
This one is definitely worth a read for phylogeneticists and phylogenomicists (is that a word?) out there: PLoS Biology: Resolving Difficult Phylogenetic Questions: Why More Sequences Are Not Enough. Philippe et al. discuss some important issues in using genomes to infer phylogenies of species in this commentary/review paper. They discuss in particular some recent studies of animal evolution but they cover a lot of useful ground here and include a good review of terminology and some of the basic issues at play. I am personally going to have to read it in more detail to help deal with some of the issues in our recent study of novel sequences in metagenomic data.
Richard Branson getting into microbial diversity (a little bit)
Just a quick one here. For those interested in the deep-sea and microbial diversity you might want to check out this article: Richard Branson launches Virgin Oceanic: deep-sea exploring submarines – Boing Boing. It discusses how Branson will be working with Katrina Edwards, Doug Bartlett and others to study microbial diversity in the deep sea as a component of his project to make dives in his personal submarine.
Request for #UCDavis affiliates to help Japan from Chancellor Katehi
Just got this email from the Chancellor of UC Davis Linda Katehi and thought I would share
To the UC Davis Community,
The crisis and devastation in Japan following the March 11 earthquake and tsunami continue, and as with any natural disaster of this magnitude, healing will be neither simple nor immediate. Our thoughts and hearts are with the Japanese people and with their many loved ones who are teaching and learning on our campus and in our community.
Inevitably, the disaster will fall from the headlines, but the UC Davis community will not look away. We will continue to reach out in comfort and with assistance to honor those who are hurting and in respect for our deep connection to Japan. Nearly 200 Japanese residents are currently studying at UC Davis, and this year, we have more than 50 Japanese faculty members visiting our campus. Our students were also at five universities in Japan at the time of the disaster, including in areas with heavy damage, and we are grateful that all are safe.
There is still much to do, and you can help, too. Please see this link for a variety of charities aiding the Japanese people:
http://www.charitynavigator.org/index.cfm?bay=content.view&cpid=1221
UC Davis has a compassionate spirit, and I thank you for proving that once again.
Linda P.B. Katehi
Chancellor
– Posted using BlogPress from my iPad
My twitter wrap up of the Joint Genome Institute User Meeting #JGIUM
Off to another meeting so don’t have time to write up details of the JGI User Meeting that just ended. But I am posting my tweets and some related tweets here. Also, apparently videos of the talks will be available soon. Will try to clean up the style of the posts ASAP but on the road …
kucsbl CSBL at Korea Univ.
RT @phylogenomics Rob Knight discussing the rationale behind his UNIFRAC metric to comparing communities using phylogeny #JGIUM
5 hours ago Favorite Retweet Reply
CackleofRad CackleofRad
@
@Tideliar @jadebio You know who else I think is awesome? Try @phylogenomics and maybe #JGIUM Not medical per se but cool things in the werks
17 hours ago
phylogenomics Jonathan Eisen
Uy vey: got home from #JGIUM & my kids had chlamydia, anthrax, E. coli, malaria, athletes foot, Helicobacter & giardia yfrog.com/gzk75auj
20 hours ago
phylogenomics Jonathan Eisen
Next up at #JGIUM Dan Distel on Shipworm symbionts – note here is a picture of Dan (foreground) from 1992 cruise http://twitpic.com/4cvb3w
22 hours ago
phylogenomics Jonathan Eisen
Apologies all – have to skip end of #JGIUM – follow this hashtag for other’s posts
23 hours ago
phylogenomics Jonathan Eisen
Scholin hacked into unsecured wireless network while on vacation in Death Valley to communicate w/ his sensors in Newport Beach #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Scholin has some of these remote sensors moored off of Newport Beach pier to survey for toxic diatoms #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Holy ecogenomic sensor batman – this remote ESP thing is very cool mbari.org/ESP/esp_2G.htm – can do DNA analysis remotely #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Next up Chris Scholin on remote detection of marine microbes in coastal waters and deep sea #JGIUM
24 Mar
phylogenomics Jonathan Eisen
In the Q and A period for Dan Distel some music has now come on over the speakers #oscars? #exitmusic #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Distel discussing proteomics of shipworm symbionts communities – uses method that focuses on proteins in symbionts not host #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Oh no – Distel mentioned the CaZome when discussing carbohydrate active enzymes in shipworm symbionts #badomicsword #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Distel: the gill symbionts are in addition to symbionts living in the gut that are known to degrade cellulose #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Distel: many (~10) types of closely related bacterial symbionts live inside the cells of shipworm in the gill #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Note – here is a link to my #PLoSONE paper with Distel on the genome of a shipworm symbiont plosone.org/article/info:d… #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Distel: shipworms are very diverse and can live off all sorts of wood & wood like stuff – major wood consuming organisms in ocean #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Dan Distel Works at the Ocean Genome Legacy foundation #JGIUM #coolgroup oglf.org/DistelCV.htm
24 Mar
phylogenomics Jonathan Eisen
Distel: shipworms (which are actually clams) cause billions of dollars of economic damage each year #JGIUM
24 Mar
doe_jgi Joint Genome Inst.
RT @phylogenomics Distel expressing thanks to JGI b/c despite claims by many that sequencing is free, nobody has told his acctg dept #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Distel expressing thanks to JGI b/c despite claims by many that sequencing is free, nobody has told his accounting department #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Next at #JGIUM Dan Distel on shipworm symbionts – note here’s a pic of Dan (foreground) from ’02 http://www.scancafe.com/p-59386415-38fa9f
24 Mar
sharmanedit Anna Sharman
Wow. MT @phylogenomics [Ed] Buckler: any two corn [maize] plants are as different from each other as humans and chimpanzees #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Next at #JGIUM Mary Ann Moran; Note paper I have w/ her is one Nature fails to make free despite promises nature.com/nature/journal… #opengate
24 Mar
phylogenomics Jonathan Eisen
Buckler described Genotyping by sequencing method from his in press #PLoSOne paper #JGIUM maizegenetics.net/images/stories…
24 Mar
phylogenomics Jonathan Eisen
Buckler suggests the genetic diversity in maize allows it to adapt to local environments better than other species #JGIUM #notbuyingit
24 Mar
phylogenomics Jonathan Eisen
Buckler: genomic domestication analysis w/ Ross-Ibarra lab from #ucdavis: little loss in diversity from landraces -> improved lines #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Buckler: Tripsacum genome has different retrotransposons than maize but otherwise may be useful as source of genetic variants #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Buckler also sequencing Tripsacum – sister genus of maize #JGIUM no chromosomal duplications, very similar gene content to maize
24 Mar
phylogenomics Jonathan Eisen
Buckler: doing maize HAPMAP2 to survey genetic diversity in corn #JGIUM
24 Mar
leonidkruglyak Leonid Kruglyak
So are yeast strains RT @phylogenomics: Buckler: any two corn plants as different as human and chimp #JGIUM #myspeciesisbetterthanyours
24 Mar
phylogenomics Jonathan Eisen
Buckler: any two corn plants are as different from each other as humans and chimpanzees #JGIUM #myspeciesisbetterthanyours
24 Mar
leonidkruglyak Leonid Kruglyak
Ed Buckler RT @phylogenomics: Next up Ed Buckley from Cornell discussing sequencing/using maize genome #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Next up Ed Buckley from Cornell discussing sequencing/using maize genome #JGIUM
24 Mar
sarahcpwilliams sarahcpwilliams
@phylogenomics enjoying yr tweets from #jgium. i’m a big fan of ley and knight. covered their work last year for hhmi: http://bit.ly/g2NEug
24 Mar
phylogenomics Jonathan Eisen
Ley : to study diversity of microbes associated with maize had to get primers that did not amplify chloroplast rDNA #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Ley: doing a QTL experiment on maize/corn treating microbes as their quantitative trait #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Ley now expanding human microbiome GWAS twin study to include surveying microbes all over body #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Ruth Ley: GWAS studies of human twins has IDd many loci that appear to affect microbial diversity – including some immune system loci #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Ruth Ley is now doing GWAS studies w/ human twins where the phenotype they are looking at is microbial diversity #JGIUM #verycool
24 Mar
phylogenomics Jonathan Eisen
Ruth Ley discussing survey of microbes in one child over two years #JGIUM
24 Mar
phylogenomics Jonathan Eisen
Ruth Ley now up at the JGI User Meeting discussing maize, human microbiotas #JGIUM … Note – I love her work #brilliant
24 Mar
kevinswilson66 Kevin Scott Wilson
@
@phylogenomics : Many thanks for your notes on #JGIUM . I was captivated by them
23 Mar
Symbiologica Juliana Mastronunzio
Thanks for tweets on the JGI meeting #JGIUM from @phylogenomics and @iGenomics.
23 Mar
sdaxen Seth D. Axen
RT @phylogenomics: Schuster: “I would like to finish my talk by discussing sequencing the devil” #JGIUM
23 Mar
Pathh1 Pat Heslop-Harrison
Done that – I found the devil in the detail. RT @phylogenomics: Schuster “like to finish my talk by discussing sequencing the devil” #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Schuster: “I would like to finish my talk by discussing sequencing the devil” #JGIUM
23 Mar
iGenomics Dawei Lin
RT @phylogenomics: Schuster: Stays away from traditional sources of ancient DNA like bone and uses hair #JGIUM
23 Mar
doe_jgi Joint Genome Inst.
RT @phylogenomics Schuster: got .1g of hair from 200 year old mammoth sample from Russia and can get mitochondrial genome #JGIUM #fb
23 Mar
phylogenomics Jonathan Eisen
Schuster: got .1g of hair from 200 year old mammoth sample from Russia and can get mitochondrial genome #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Schuster working on thylacine (tasmanian tiger) mitochondrial genomes #JGIUM thylacine.psu.edu
23 Mar
phylogenomics Jonathan Eisen
For more on mammoth genomics see mammoth.psu.edu #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Schuster: when you sample extinct organisms you have to remember that different samples may come from different times #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Schuster: “You would not believe how much mammoth hair I have washed off myself” #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Schuster: Stays away from traditional sources of ancient DNA like bone and uses hair #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Schuster – redundancy in genome sequencing with ancient genomes helps build quality genome assemblies #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Schuster: one reason to focus on mitochondrial genomes is that there are lots of copies of the genome per cell #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Schuster : “dont forget about mitochondrial genomes” still lots of species that do not have mt genome sequences #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Schuster: discussing mammoths, moas, thylacines, tasmanian devils and polar bears #JGIUM #museomics #conservation #endangered
23 Mar
phylogenomics Jonathan Eisen
For more on Schusters work on extinct species see cidd.psu.edu/people/scs19 #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Next up Stephan Schuster discussing the Genomics of Extinct and Endangered Species #JGIUM #museomics
23 Mar
Energy_Science Energy Science News
@doe_jgi: RT @phylogenomics Why #badomics words can also be very good: a case in study with museomics #JGIUM http://ff.im/-zyA86 #fb
23 Mar
doe_jgi Joint Genome Inst.
RT @phylogenomics Why #badomics words can also be very good: a case in study with museomics #JGIUM http://ff.im/-zyA86 #fb
23 Mar
phylogenomics Jonathan Eisen
Why #badomics words can also be very good: a case in study with museomics #JGIUM http://ff.im/-zyA86
23 Mar
phylogenomics Jonathan Eisen
Eddy Rubin at the #JGIUM is soliciting input from crowd on future needs of the community
23 Mar
Energy_Science Energy Science News
@doe_jgi: #JGIUM bingo anyone? DOE JGI Director Rubin mentions @phylogenomics GEBA project in his talk on future of DOE JGI #fb
23 Mar
phylogenomics Jonathan Eisen
The perils of giving out #badomics word awards – a prior recipient at #JGIUM just told me he’s still angry at me phylogenomics.blogspot.com/2009/01/worst-…
23 Mar
doe_jgi Joint Genome Inst.
#JGIUM bingo anyone? DOE JGI Director Rubin mentions @phylogenomics GEBA project in his talk on future of DOE JGI #fb
23 Mar
Energy_Science Energy Science News
@doe_jgi: #JGIUM Rob Knight: “There is one universal tree of life which is why projects such as @phylogenomics GEBA are so critical” #fb
23 Mar
phylogenomics Jonathan Eisen
All I can say is that when I was rejected by HHMI a few yrs ago I felt better when I heard Rob Knight got it b/c, well, he rocks #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Rob Knight – in human microbiome studies you actually need VERY few sequences per sample to get overall trends #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Rob Knight: “There is one universal tree of life” and giving props to my GEBA genomic encyclopedia project #JGIUM
23 Mar
doe_jgi Joint Genome Inst.
#JGIUM Rob Knight: “There is one universal tree of life which is why projects such as @phylogenomics GEBA are so critical” #fb
23 Mar
phylogenomics Jonathan Eisen
Rob Knight discussing the rationale behind his UNIFRAC metric to comparing communities using phylogeny #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Rob Knight discussing how sequencing has gotten so cheap and high throughout that analysis tools are the limiting step in many cases #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Rob Knight now up at #JGIUM – he publishes more cool papers per month than just about anyone in microbial research
23 Mar
phylogenomics Jonathan Eisen
Silver: took sugar secreting cyanobacterium and got macrophage to take them up and they survive a little bit #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Silver: injected sugar secreted cyanobacterium into zebrafish zygotes and get functional fish with cyanos all throughout them #jgium
23 Mar
phylogenomics Jonathan Eisen
Silver: engineered a cyanobacterium secrete sugars so thought maybe they could use this to make photosynthetic animals #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Silver also interested in biohydrogen production #JGIUM but two problems: most hydrogenases are oxygen sensitive and electron competition
23 Mar
phylogenomics Jonathan Eisen
Silver is working on engineering 3hydroxypropionate carbon fixation pathway from Chloroflexus in E. Coli #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Silver claimed that Cyanobacteria are responsible for 50% of photosynthesis on earth but I think that must be too high #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Silver working on redesigning photosynthesis via cyanobacteria #JGIUM – says they need to learn a lot of biology still
23 Mar
phylogenomics Jonathan Eisen
Silver: though she tries to get $$ from basic scion agencies , they never fund her #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Pam Silver: uses “redesign of a system can test our understanding of it’s components” to try to get $$ from basic science agencies #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Pam Silver “Biology is the technology of this century” is the message she wants to gt across #JGIUM
23 Mar
phylogenomics Jonathan Eisen
Jerry Tuskan getting some hard but good questions after his talk at #JGIUM – Q and A much more interesting than talks usually
23 Mar
phylogenomics Jonathan Eisen
Next up at #JGIUM Pam Silver – not only brilliant – but also her lab is the source of a good Lady Gaga spoof youtube.com/watch?v=ZilqYp…
23 Mar
phylogenomics Jonathan Eisen
Tuskan using Genome and RNA sequencing and high throughout phenotyping for massive poplar association study #jgium
23 Mar
phylogenomics Jonathan Eisen
At #JGIUM listening to Jerry Tuscan discuss poplar genomics – the place is packed yfrog.com/h82u1nnj
23 Mar
iGenomics Dawei Lin
@
@phylogenomics Terry Hazen talked about it. It can be easily forgot how many people worked behind the scene. #jgium
22 Mar
phylogenomics Jonathan Eisen
Tell me about it: “@iGenomics: It is hard to include all people working on a project into the author list these days. #jgium”
22 Mar
Energy_Science Energy Science News
@doe_jgi: RT @phylogenomics: Personal trivia re: SLAC director Perisis Drell at #JGIUM – previous director Artie Bienenstock was a st…
22 Mar
phylogenomics Jonathan Eisen
Personal trivia re: SLAC director Perisis Drell at #JGIUM – previous director Artie Bienenstock was a student of my grandfather Ben Post
22 Mar
EpiExperts Epigenetics Experts
RT @phylogenomics @doe_jgi: SLAC National Accelerator Lab Director Persis Drell kicks off DOE JGI User Mtg 5pm-use #JGIUM to follow
22 Mar Favorite Retweet Reply
Why #badomics words can also be very good: a case in study with museomics #JGIUM
Well, my snarky blog style with some of my awards comes with some risks. Today I experienced one of them. Stephan Schuster gave me some serious major grief over a post I wrote a few years ago. In the post I gave him an award for “Worst new omics word” for the word museomics. I gave this award because, well, I don’t like the word. I stand by my complaints about the word. But Stephan did highlight ( in his comments to me in the back of the room at the JGI User Meeting) that the word has been remarkably useful at getting money and attention for museum based genomics studies.
So I guess here is the dilemma. I realize new omics words can get attention. Omics is after all very hot still. But I write about #badomics words both because I think it is fun and also because I think people are careless with the language much of the time. Many omics words are really awful with no benefits. But some, fit my criteria as being badomics words, but can have positive benefits. To the public, the word museomics probably conjures up exactly what museum people want – the image of cutting edge science. Though I love museums, to some the conjure up images of dust and cobwebs and crotchety old scientists wandering around the halls in the dark. So the term museomics may indeed turn this on it’s head. This term even to me conjures up images of museums doing cool things. So alas, I guess this is a mea culpable of sorts. I still think the word itself is not great. After all, if you think about it, all it really means is doing genomics with museum specimens. But clearly Stephan has a good point … Getting attention to science is a good thing. So I guess badomics words can be used for good.
I am still going to snark about badomics words. But I will try to make more mention of the potential value they have in spreading science and omics to the masses ….
– Posted using BlogPress from my iPad
UPDATE 5/13/2013
What a wimp out. Museomics is such a bad bad bad omics word.