Hello Everyone,
I have a few updates with regard to the Undergraduate Research Conference.
The abstract that I turned in originally had a few mistakes, so I made sure to contact the Undergraduate Research office at the Student Community Center. Initially, they said that they wouldn’t allow any corrections upon submission of the abstract, but I was slightly insistent, so they agreed.
The updated abstract has been developed after considering everyone’s inputs and modifying the original abstract as necessary. As of now, it reads as below:
Aquarium Biogeography and Succession of Microbial Communities in Aquatic Environments
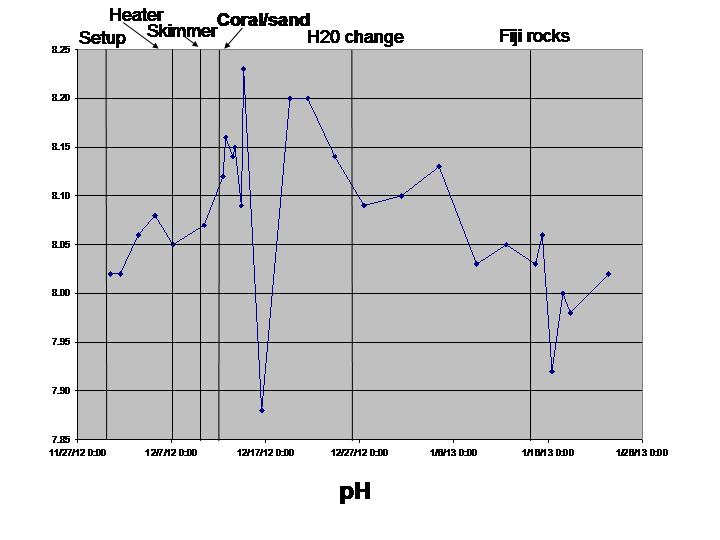
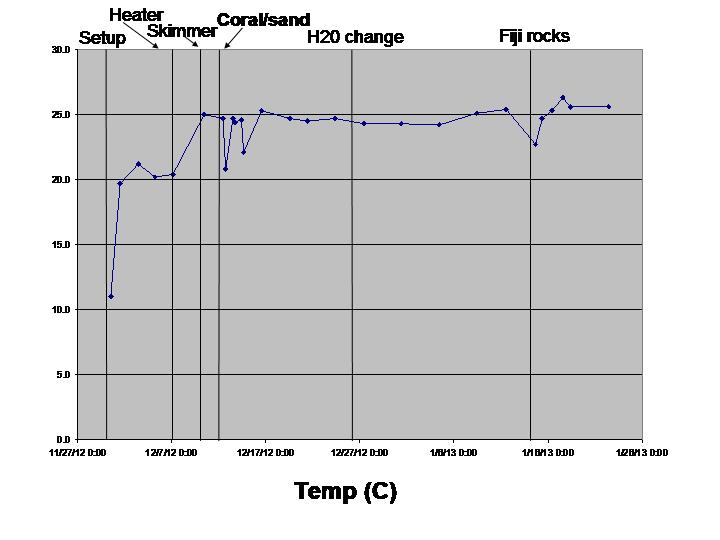
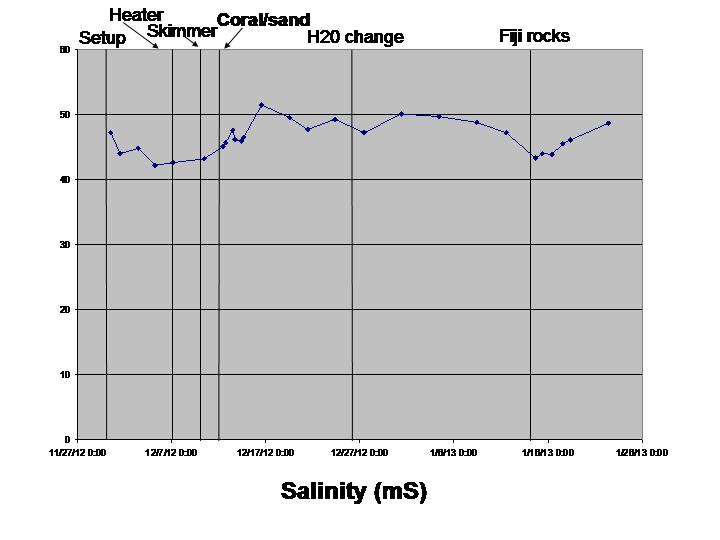
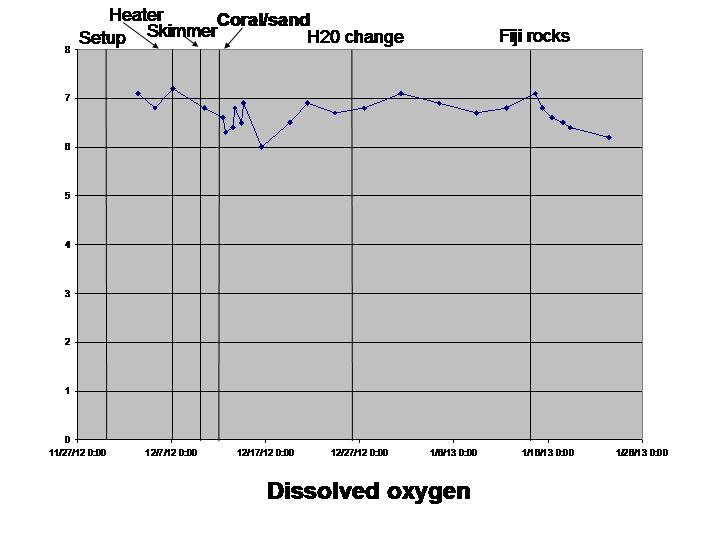
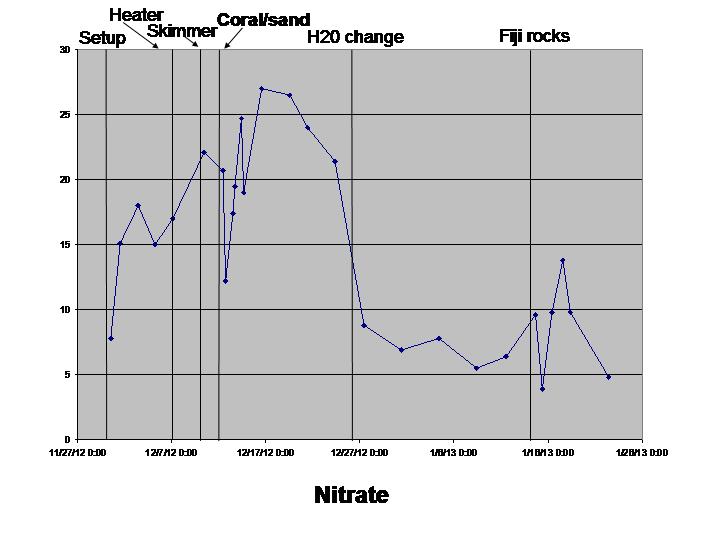
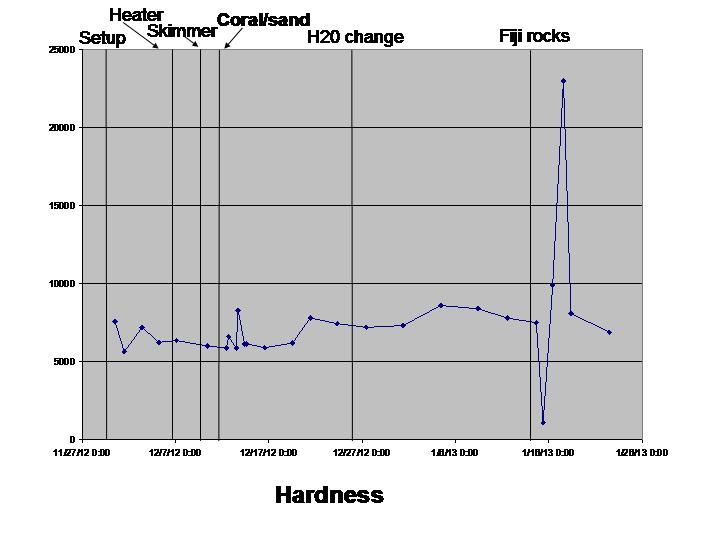
The biological sciences teaching laboratory for UC Davis maintains a wide variety of large aquaria, including freshwater and marine tanks, designed to mimic both temperate and tropical ecosystems. In this study, we seek to better understand the biogeography of microbial communities associated with these tanks. Using a culture-independent, DNA-based community census approach (i.e., 16S rRNA PCR surveys), we tackle two main questions: 1) how do microbial communities vary with respect to various environmental parameters (temperature, salinity, pH, oxygen and nitrogen concentrations) both within and across tanks; and 2) how does the structure of microbial communities change in response to ecosystem disturbance. To address this second question, we coincided our data collection with the establishment of two new aquarium systems (coral ponds). We use high-throughput DNA sequencing of the 16S rRNA gene to generate a microbial community profile from hundreds of samples. These microbial community profiles are then compared across tanks (marine vs. fresh water, warm vs. cold) and in response to disturbances introduced during the establishment of the coral ponds.
I hope this acts as an almost accurate summary of what we are doing. The abstract was accepted, reviewed and approved by the URC people, so we are all set as far as that is concerned.
I need to also come up with a poster to represent our lab and the kind of work we do, and I get access to free poster printing services for that, so yay! This work is going to start next quarter, and I’m hoping that some of you will pitch in and help me out to make a good job of it and represent our lab in the way it deserves to be showcased. Also, I don’t know if others are allowed to co-present with me. If you are interested, do contact the URC office, I think it’s a great opportunity.
I attended the mandatory meeting for all participants of the conference. It was hosted by Tammy Hoyer of the Undergraduate Research Office today. It was pretty informative in the sense it gave me information on all the resources that were made available to me and what I could expect at the Undergraduate Research Conference.
Also, Tammy mentioned about the Undergraduate Research Journal, Explorations. They are currently taking papers, and I believe the deadline is 6th June, if I heard correctly. I know we wouldn’t have completed all of our research by then, but just letting you know of the opportunity.
Well, that’s all for now.
Also, starting next quarter, I will try my best to do the 8 hour a week thing, will be coming in on Mondays and Fridays, a 4+4. I hope to work with most of you during that time.
Good luck for finals!