Category: Misc.
Special Guest Post & Discussion Invitation from Matthew Hahn on Ortholog Conjecture Paper
I am very excited about today’s post. It is the first in what I hope will be many – posts from authors of interesting papers describing the “Story behind the paper“. I write extensive detailed posts about my papers and also have tried to interview others about their papers if they are relevant to this blog. But Matthew Hahn approached me recently about the possibility of him writing up some details on his recent paper on the functions of orthologs vs. paralogs. So I said “sure” and set up a guest account for him to write up his comments and details of the paper.
For those of you who do not know, Matt is on the faculty at U. Indiana. He was a post doc at UC Davis so I have a particular bias in favor of him. But his recent paper has generated some controversy (I posted some links about it here). So it is great to get some more detail from him. In addition, I note, I am also using this approach to try and teach people how easy it is to write a blog post by getting them guest accounts on Blogger and letting them write up something with links, pictures, etc. So hopefully we can get more scientists blogging too.
Anyway – without any further ado – here is Matt’s post:
Following Jonathan’s excellent example of how explaining the history of a project helps to illuminate how the process of science actually happens, I thought I’d start by giving a bit of history behind our study, and the paper that we recently published in PLoS Computational Biology (http://www.ploscompbiol.org/article/info:doi/10.1371/journal.pcbi.1002073). And then I’ll address the critics…
—————————
Note – Toni Gabaldón was trying to post a detailed response but Blogger kept cutting him off with a character limit. So I have posted his response below.
I appreciate the effort by Matthew Hahnn on explaining the story behind his paper on the so-called “Ortholog conjecture” and on facing some of the criticism. This paper attracted my interest as that of many others that work on or just use orthology. For instance it was chosen by one of my postdocs for our “Journal Club” meeting. And it was discussed during our last “Quest for Orthologs” meeting in Cambridge. I think is raising a necessary discussion and therefore I think is a good paper. This does not mean that I fully agree with the interpretation and conclusions ;-). I hope to modestly contribute to this debate with the following post.
I think one of the causes that this paper has caused so much debate is that the conclusions seem to challenge common practice (inferring function from orthologs), and could be interpreted as the need of changing the strategies of genome annotation. I think, however, that one should interpret carefully these results before start annotating based on paralogous proteins. As I will discuss below one of the problems is that we need to agree in what is the conjecture to then agree in how to test it. I see three main points that can be a source of confusion: i) the issue of what is actually stated by this conjecture, ii) the issue of annotation, and iii) the issue of time
1) What is the “ortholog conjecture”?
Or in other terms, when should we expect orthologs to be more likely to share function than paralogs?. Always? Of course not. All of us would agree that two recently duplicated paralogs are likely to be more similar in function than two distant orthologs, so it is obvious that the conjecture is not simply “orthologs are more similar in function than paralogs”. In reality the expectation that orthologs are more likely to be similar in function than paralogs, as least this is how I interpret it, is directly related to the effect that duplication have on functional divergence. If gene duplication has some effect on functional divergence (even in not 100% of the cases), then, given all other things equal (divergence time, story of speciation/duplication events – except fpr the duplication defining the orthologs) one would expect orthologs to be more likely to conserve function.
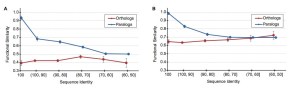
I think this complexity is not well considered (by many authors, in general). Hahn refeers to the famous review of orthology by Koonin (2005) as the source for the term “ortholog conjecture”. However, In that paper this conjecture is discussed always within the context of genes accross two particular species, whether in Hahn’s paper it is taken as well to other contexts. Thus, the proper context in which to test this conjecture is only between orthologs and between-species paralogs. As we can see, Red and purple lines in Hahn paper in figure2 do not show any clear difference.
Secondly, Koonin was very cautions in his paper, stating that he was referring to “equivalent functions” and not exactly the same “function”, correctly implying that the functional contexts would be different in the two different species. This brings me to the next point.
ii) annotation
If the expectation of functional conservation of orthologs refers to a given pair of species, then it makes no sense to test that expectation between paralogs within the same species and orthologs in different species. We were interested in this issue and it took us some effort to control for this “species” influence on the comparison, if you are interested you can read our paper on divergence of expression profiles between orthologs and paralogs (http://www.ncbi.nlm.nih.gov/pubmed/21515902)
As Hahn founds, and it was anticipated by Koonin in that review, there is a huge influence of the “species context”, a big constraint of what fraction of the function is shared. Indeed I think is the dominant signal in Hahn’s paper. Why is that? One possibility is that the functional context determines the function, I agree. However, we should not discard biases in how different communities working around a model species define processes and function, also the type of experiments that are usually done. For instance experimental inference from KO mutants might be common from mouse, but I guess is not the case in humans (!!). I think this may be having a big influence and might even be the dominant signal in Hahns paper.
Finally function has many levels and I expect subfunctionalization mostly affect lower levels (i.e. more specific). Biases may also
exist in the level of annotation between species or between families of different size (contributing more or less to the orthologs/paralogs class).
Microarray data are less likely to be subject to biases (although some may exist), at least they should be expected to be free of “human interpretation biases” and so Hahn and colleaguies did well, in my opinion, of testing that dataset. It is important to note that for microarrays and for orthologs and between-species paralogs (which I think is the right frame for testing the conjecture) ortholgs are more likely to share an expression context. This is compatible to what we found in the paper mentioned above, and compatible with the orthology conjecture as stated by koonin (accross species)
iii) time
Finally, one aspect which I think is fundamental is the notion of “divergence time”. Since paralogs can emerge at different time-scales they are composed by a heterogeneous set of protein pairs. Most of comparisons of orthologs and paralogs (Hahn’s as well) use sequence divergence as a proxy of time. However this is only a poor estimate, specially when duplications (as in here) are involved (we explored this issue in the past: http://www.ncbi.nlm.nih.gov/pubmed/21075746). This means that for a given divergence time paralogs may have larger sequence divergence than orthologs at the same divergence time, or otherwise (if gene conversion is playing a role). Is the conjecture based on sequence divergence or on divergence time?, I think the initial sense of using orthology to annotate accross species is based on the notion of comparing things at the same evolutionary distance. Thus basing our conclusions on divergence times might not be the proper way of doing it.
CONCLUSIONS AND PROPOSAL FOR RE-STATEMENT
To conclude, and with the intention of going beyond this particular paper,
I would finish by saying that the key to the problem lies on how we interpret the so-called “ortholog conjecture” or how are our expectations on how function evolves. What I get from re-reading Eugene Koonin’s paper and how I am using that “assumption” in my day-to-day work is the following:
“Orthologs in two given species are more likely to share equivalent functions than paralogs between these two species”
Therefore the notion of “accross the same pair of species” is important and thus only part of the comparisons made by Hahn and colleagues could directly test this. Looking at the microarray and between-species comparisons data, the conjecture may even hold true!!
I, however, do think that the conjecture as stated above is limited and does not capture the complexity of orthology relationships. Indeed us, and many other researchers, are tuning the confidence of the orthology-based annotation based on whether the orthologs are one-to-one, one-to-many or many-to-many, even when orthologs are “super-orthologs” (with no duplication event in the lineages separating the two orthologs).
Since, the underlying assumption of the ortholog conjecture is that duplication may (not necessarily always) promote functional shifts, then many-to-many orthology relationships will tend to include orthologous pairs with different functions.
Thus I would re-state the conjecture (or expectation) as follows:
“In the absence of additional duplication events in the lineages separating them, two orthologous genes from two given species are more likely to share equivalent functions than two paralogs between these two species”
This would be a more conservative expectation, which is closer to the current use of orthology-based annotation that tends to identify one-to-one orthologs, rather than any type.
When duplications start appearing in subsequent lineages thus creating one- or many-to-many orthology relationships, the situation is less clear. Following the assumption that duplications may promote functional divergence. Then one could expand the conjecture by “the more duplications in the evolutionary history separating two genes, the lower the expectation that these two genes would share equivalent functions”.
I wrote this contribution on the fly, and surely there are ways of expressing this in more appropriate terms. In any case I hope I made clear the idea that the conjecture emerges from the notion of duplications causing functional shifts and that our expectations will be clearer if expressed on those terms. This goes on the lines of what Jonathan Eisen mentioned on considering the whole phylogenetic story to annotate genes.
Under this perspective, the real important hypothesis is that “duplications tend promote functional shifts”, I think this is based on solid grounds and has been tested intensively in the past.
Cheers,
Toni Gabaldón
Interested in sex? How about in bacteria? Then these #PLoSGenetics papers are for you
Well I was torn about this. Should I title the post ” ICE, ICE, Bacterial BABIES” or say something about sex? I settled on sex, but not sure if that was wise.
Anyway – quick post to say that there are two papers from PLoS Genetics last month that caught my eye. They are
- PLoS Genetics: The Repertoire of ICE in Prokaryotes Underscores the Unity, Diversity, and Ubiquity of Conjugation. Guglielmini J, Quintais L, Garcillán-Barcia MP, de la Cruz F, Rocha EPC (2011) PLoS Genet 7(8): e1002222. doi:10.1371/journal.pgen.1002222
- PLoS Genetics: A Broad Brush, Global Overview of Bacterial Sexuality Achtman M (2011) PLoS Genet 7(8): e1002255. doi:10.1371/journal.pgen.1002255
The latter is a “review” paper linked to the first one which is a research paper. The papers together provide both a good background and a window into modern studies of “ICEs” or integrative conjugative elements in bacteria.
I like the summary from the first paper:
Some mobile genetic elements spread genetic information horizontally between prokaryotes by conjugation, a mechanism by which DNA is transferred directly from one cell to the other. Among the processes allowing genetic transfer between cells, conjugation is the one allowing the simultaneous transfer of larger amounts of DNA and between the least related cells. As such, conjugative systems are key players in horizontal transfer, including the transfer of antibiotic resistance to and between many human pathogens. Conjugative systems are encoded both in plasmids and in chromosomes. The latter are called Integrative Conjugative Elements (ICE); and their number, identity, and mechanism of conjugation were poorly known. We have developed an approach to identify and characterize these elements and found more ICEs than conjugative plasmids in genomes. While both ICEs and plasmids use similar conjugative systems, there are remarkable preferences for some systems in some elements. Our evolutionary analysis shows that plasmid conjugative systems have often given rise to ICEs and vice versa. Therefore, ICEs and conjugative plasmids should be regarded as one and the same, the differences in their means of existence in cells probably the result of different requirements for stabilization and/or transmissibility of the genetic information they contain.
That should be enough to get people started. And that is alas all I have time to write about here.
Put down what you are doing & read this article: Amy Harmon "Autistic & seeking a place in an adult world"
Assemblathon 1 paper out, includes many #UCDavis folks, though @vsbuffalo name backwards
Can I just say I love Biomed Central #OpenAccess
Well, I have given Biomed Central a bit of snarky grief the last few days over a few things. First, I posted about to my Posterous site (but not here) a little comment about how their web site looks weird in safari:
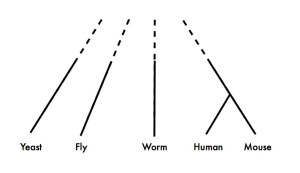
Then I posted to this blog a little ditty about how I did not like some parts of a phylogenetic tree they use in marketing:No award to give out but here are some lessons in using Google’s image search to find an image source
My main complaint was the poor treatment of microbes in the tree. In that post I discussed how I used google image search to trace the tree
to a few sites and discovered that they recognized it was a bit of a biased tree. And I noted they had fridge magnets that had the tree and how I wanted one.
And, well, they have responded brilliantly.
Matthew Cockerill posted to my posterous site about how he was looking into the Safari issue and then, they fixed it (it was a font display issue).
And then today in the mail I received a gift and a note
The note reads “”We’ll do justice to the microbial world one day”.
Indeed.
I note, even without their responses, I truly love Biomed Central. I published my first open access paper was published in a Biomed Central Journal, Genome Biology: http://genomebiology.com/2000/1/6/research/0011 and I have published quite a few articles in their journals including:
- Introducing W.A.T.E.R.S.: a Workflow for the Alignment, Taxonomy, and Ecology of Ribosomal Sequences
- Acidithiobacillus ferrooxidans metabolism: from genome sequence to industrial applications
- Refined annotation and assembly of the Tetrahymena thermophila genome sequence through EST analysis, comparative genomic hybridization, and targeted gap closure
- A genomic analysis of the archaeal system Ignicoccus hospitalis-Nanoarchaeum equitans
- A simple, fast, and accurate method of phylogenomic inference
- New evolutionary frontiers from unusual virus genomes
Biomed Central was THE pioneer for truly open access publications in biology and they are still doing great things. I note in addition, they do a very good job covering microbiology not only in their general journals but also with specific microbiology focused journals including:
- Annals of Clinical Microbiology and Antimicrobials
- Antimicrobial Resistance and Infection Control
- BMC Infectious Diseases
- BMC Microbiology
- Gut Pathogens
- Microbial Cell Factories
- Microbial Informatics and Experimentation
- Virology Journal
So it seems – you are already doing some justice to the microbial world.
C-DEBI Research Support > Request for Research Proposals
 |
| Katrina Edwards on the Atlantis |
I have always been fascinated by life in extreme places on the planet. And somehow I have managed to do projects on microbes from places like Antarctica, boiling hotsprings in Yellowstone and Kamchatka, acid pools, and more. The extremes are fascinating to me because they tell us a lot about the limits of life as well as indirectly about life in “normal” places.
And of course, I am not alone. Many many scientists are fascinated by life’s extremes. But not everyone ends up studying life in extreme environments of course. One reason for this is that many extreme environments that might be of interest are kind of hard to study. Consider the deep sea. Not so easy to do work there and just getting samples can be a massive undertaking.
Just imagine though. What if there were a way to “tag along” on an existing project studying life’s extremes at no cost to you or your grants? Even better what if there were a way to get extra funds to not just tag along on a project but to carry out detailed research at the same time?
Well, amazingly, there is such a chance right now. The C-DEBI “Center for Dark Energy Biosphere” project is calling for proposals. C-DEBI Research Support > Request for Research Proposals
They have money. They have drills. They have been and will continue to be collecting lots of samples from the bottom of the ocean and the crust below. They are doing a bunch of microbiology (as well as other things). And they are calling for people out there to join them in various ways including;
- Research Proposals
- Research and Travel Exchange Program
- Post doc scholar program
- Graduate fellow program
- Education and outreach proposals
- http://www.darkenergybiosphere.org/return-to-northpond/
- http://aam.darkenergybiosphere.org/
- http://www.darkenergybiosphere.org/classroomconnection
- http://joidesresolution.org/node/1983
I note – I was a visiting scientist for a few days at one of the C-DEBI meetings about evolution earlier this year. It was a great meeting – on Catalina Island – and I wrote a VERY long blog post about it: The Tree of Life: A “work” trip to Catalina Island: USC, Wrigley, C-DEBI, dark energy biosphere, Virgin Oceanic, Deep Five, & more. You can learn more about the C-DEBI project by reading that post. And you can look at my pretty pictures below:
I note in addition, I am forever in debt to Katrina Edwards the PI of the C-DEBI project ever since she gave a frigging awesome tour to my kids of the Atlantis when it was docked in San Francisco
But regardless of the personal connections I have to C-DEBI, the project is very interesting and the fact that they are offering up funds to support “outsiders” who want to participate in the project in some way is great.
Great paper showing the potential power of comparative and evolutionary genomics in #PLoS Genetics
There is a wonderful paper that has just appeared in PLoS Genetics I want to call people’s attention to: PLoS Genetics: Emergence and Modular Evolution of a Novel Motility Machinery in Bacteria
In the paper, researchers from CNRS and Aix-Marseille in France used some nice comparative and evolutionary genomics analyses along with experimental work to characterize the function and evolution of gliding motility in bacteria.
Their summary of their work:
Motility over solid surfaces (gliding) is an important bacterial mechanism that allows complex social behaviours and pathogenesis. Conflicting models have been suggested to explain this locomotion in the deltaproteobacterium Myxococcus xanthus: propulsion by polymer secretion at the rear of the cells as opposed to energized nano-machines distributed along the cell body. However, in absence of characterized molecular machinery, the exact mechanism of gliding could not be resolved despite several decades of research. In this study, using a combination of experimental and computational approaches, we showed for the first time that the motility machinery is composed of large macromolecular assemblies periodically distributed along the cell envelope. Furthermore, the data suggest that the motility machinery derived from an ancient gene cluster also found in several non-gliding bacterial lineages. Intriguingly, we find that most of the components of the gliding machinery are closely related to a sporulation system, suggesting unsuspected links between these two apparently distinct biological processes. Our findings now pave the way for the first molecular studies of a long mysterious motility mechanism.
Basically, they started with some genetic and functional studies in Myxococcus xanthus. They analyzed these in the context of the genome sequence (note – I was a co-author on the original genome paper). And then they did some extensive comparative and evolutionary analysis of these genes, producing some wonderful figures along the way such as:
Based on their analysis they then came up with some hypotheses as to which genes were involved in key parts of gliding motility and what their biochemical functions were and they then went and confirmed this with experiments. I am not going to go into detail on the functional work they did but you can read their paper for more details.
They wrapped up their paper by proposing an model for the evolutionary history of gliding motility. I am not sure I buy all components of their model since our sampling of genomes right now is still very poor, but they have a pretty detailed theory captured in part in this figure:
Anyway – I don’t have much time right now to provide more detail on the paper. But it is definitely worth checking out.
Storification of my notes/tweets from #UCDavis CLIMB Symposium "The infant gut microbiome: prebiotics, probiotics and establishment"
I made a Storify posting for the CLIMB Symposium I participated in yesterday. First I am reposting my summary of what the symposium was about which I posted the day before the meeting:
There is a symposium tomorrow at UC Davis organized by a undergraduates in the CLIMB program. CLIMB stands for “Collaborative Learning at the Interface of Mathematics and Biology (CLIMB)” and is a program that emphasizes hands-on training using mathematics and computation to answer state-of-the-art questions in biology. A select group of undergraduates participate in the program and this summer the students had to do some sort of modelling project. Somehow I managed to convince them to do work on human gut microbes. And they have done a remarkable job.
As part of their summer work, they organized a symposium on the topic and their symposium takes place tomorrow. Details are below.
The Infant Gut Microbiome: Prebiotics, Probiotics, & Establishment
- Jonathan Eisen, UC Davis “DNA and the hidden world of microbes”
- Mark Underwood, UC Davis “Dysbiosis and necrotizing enterocolitis”
- Ruth Ley, Cornell University “Host-microbial interactions and metabolic syndrome”
- CLIMB 2010 cohort “Breast milk metabolism and bacterial coexistence in the infant microbiome”
- David Relman, Stanford University “Early days: assembly of the human gut microbiome during childhood”
- Bruce German, UC Davis
The only major issue for me is I am losing my voice. So we will see how this goes. Though I note I have gotten some very sage advice on how to treat my voice problem via the magic of twitter. If I do not collapse I will also be tweeting/posting about the other talks during the day.
Anyway – here is the storification:
http://storify.com/phylogenomics/climb-symposium-at-uc-davis.js<a href=”http://storify.com/phylogenomics/climb-symposium-at-uc-davis” target=”_blank”>View “CLIMB Symposium at UC Davis” on Storify</a>
Coming Monday at #UCDavis "The Infant Gut Microbiome: Prebiotics, Probiotics, & Establishment"
Just a little announcement here. There is a symposium tomorrow at UC Davis organized by a undergraduates in the CLIMB program. CLIMB stands for “Collaborative Learning at the Interface of Mathematics and Biology (CLIMB)” and is a program that emphasizes hands-on training using mathematics and computation to answer state-of-the-art questions in biology. A select group of undergraduates participate in the program and this summer the students had to do some sort of modelling project. Somehow I managed to convince them to do work on human gut microbes. And they have done a remarkable job.
As part of their summer work, they organized a symposium on the topic and their symposium takes place tomorrow. Details are below.
The Infant Gut Microbiome: Prebiotics, Probiotics, & Establishment
Monday, 12 September 2011, 9am-4pm
Life Sciences 1022
UC Davis
9:00-9:10 Introduction
9:10-9:40 Jonathan Eisen, UC Davis
“DNA and the hidden world of microbes”
9:40-10:40 Mark Underwood, UC Davis
“Dysbiosis and necrotizing enterocolitis”
10:40-10:50 break
10:50-11:50 Ruth Ley, Cornell University
“Host-microbial interactions and metabolic syndrome”
11:50-12:00 general discussion
12:00-1:00 lunch
1:00-2:00 CLIMB 2010 cohort
“Breast milk metabolism and bacterial coexistence in the infant microbiome”
2:00-2:10 break
2:10-3:10 David Relman, Stanford University
“Early days: assembly of the human gut microbiome during childhood”
3:10-3:40 Bruce German, UC Davis
3:40-4:00 next steps
The only major issue for me is I am losing my voice. So we will see how this goes. Though I note I have gotten some very sage advice on how to treat my voice problem via the magic of twitter. If I do not collapse I will also be tweeting/posting about the other talks during the day.